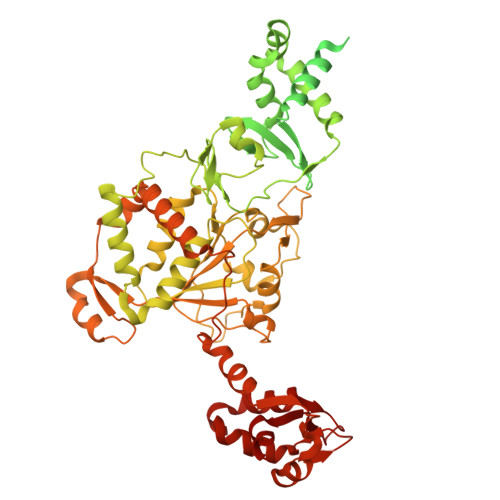
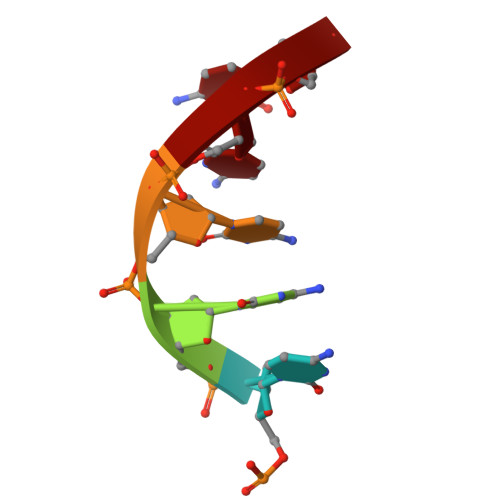
Essential and multifunctional mpox virus E5 helicase-primase in double and single hexamer.
Xu, Y., Wu, Y., Zhang, Y., Gao, K., Wu, X., Yang, Y., Li, D., Yang, B., Zhang, Z., Dong, C.(2024) Sci Adv 10: eadl1150-eadl1150
- PubMed: 39167653 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adl1150
- Primary Citation Related Structures:
8WGY, 8WGZ, 8WH0, 8WH2, 8WH3, 8WH4, 8WH6 - PubMed Abstract:
An outbreak of mpox virus in May 2022 has spread over 110 nonpandemic regions in the world, posing a great threat to global health. Mpox virus E5, a helicase-primase, plays an essential role in DNA replication, but the molecular mechanisms are elusive. Here, we report seven structures of mpox virus E5 in a double hexamer (DH) and six in single hexamer in different conformations, indicating a rotation mechanism for helicase and a coupling action for primase. The DH is formed through the interface of zinc-binding domains, and the central channel density indicates potential double-stranded DNA (dsDNA), which helps to identify dsDNA binding residues Arg 249 , Lys 286 , Lys 315 , and Lys 317 . Our work is important not only for understanding poxviral DNA replication but also for the development of novel therapeutics for serious poxviral infections including smallpox virus and mpox virus.
- Department of Thyroid and Breast Surgery, Zhongnan Hospital of Wuhan University, State Key Laboratory of Virology, School of Pharmaceutical Sciences, Wuhan University, Wuhan 430071, China.
Organizational Affiliation: