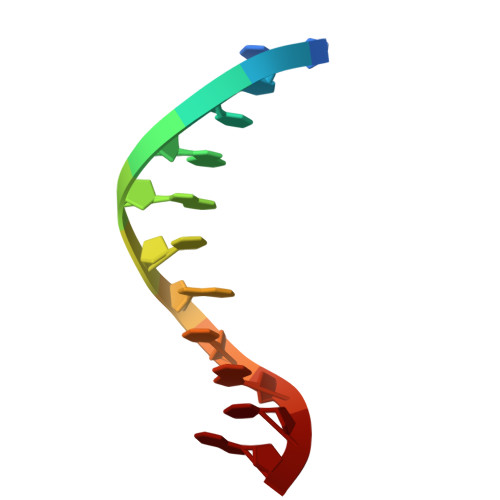
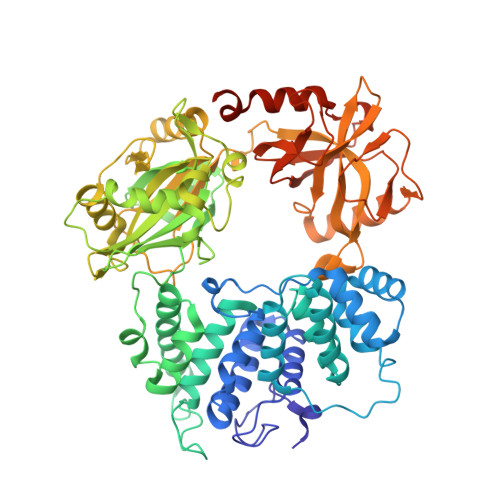
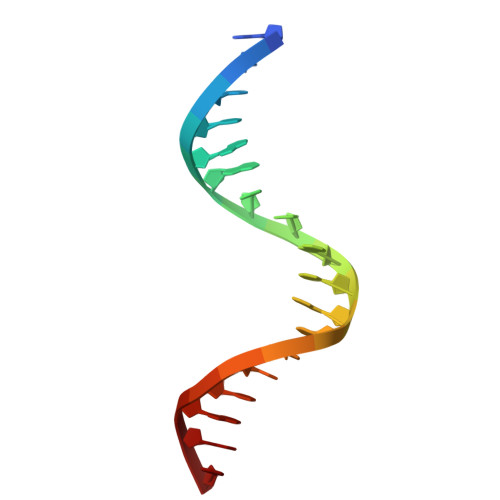
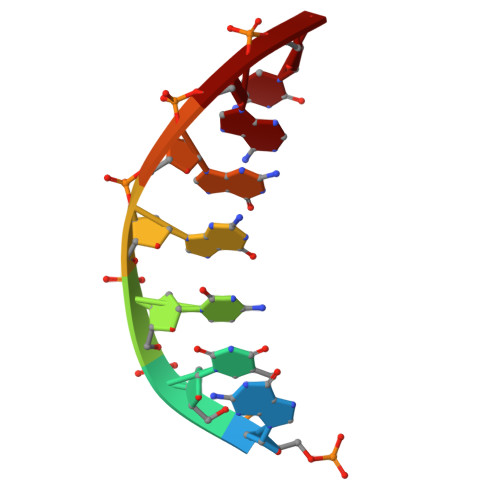
Structures of LIG1 provide a mechanistic basis for understanding a lack of sugar discrimination against a ribonucleotide at the 3'-end of nick DNA.
Balu, K.E., Gulkis, M., Almohdar, D., Caglayan, M.(2024) J Biological Chem 300: 107216-107216
- PubMed: 38522520 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2024.107216
- Primary Citation Related Structures:
8VDN, 8VDS, 8VDT, 8VZL, 8VZM - PubMed Abstract:
Human DNA ligase 1 (LIG1) is the main replicative ligase that seals Okazaki fragments during nuclear replication and finalizes DNA repair pathways by joining DNA ends of the broken strand breaks in the three steps of the ligation reaction. LIG1 can tolerate the RNA strand upstream of the nick, yet an atomic insight into the sugar discrimination mechanism by LIG1 against a ribonucleotide at the 3'-terminus of nick DNA is unknown. Here, we determined X-ray structures of LIG1/3'-RNA-DNA hybrids and captured the ligase during pre- and post-step 3 the ligation reaction. Furthermore, the overlays of 3'-rA:T and 3'-rG:C step 3 structures with step 2 structures of canonical 3'-dA:T and 3'-dG:C uncover a network of LIG1/DNA interactions through Asp570 and Arg871 side chains with 2'-OH of the ribose at nick showing a final phosphodiester bond formation and the other ligase active site residues surrounding the AMP site. Finally, we demonstrated that LIG1 can ligate the nick DNA substrates with pre-inserted 3'-ribonucleotides as efficiently as Watson-Crick base-paired ends in vitro. Together, our findings uncover a novel atomic insight into a lack of sugar discrimination by LIG1 and the impact of improper sugar on the nick sealing of ribonucleotides at the last step of DNA replication and repair.
- Department of Biochemistry and Molecular Biology, University of Florida, Gainesville, Florida, USA.
Organizational Affiliation: