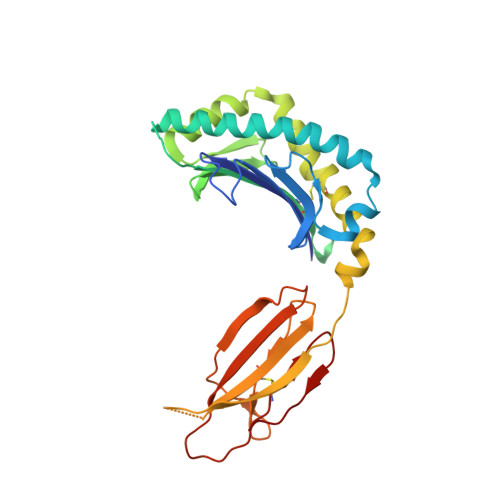
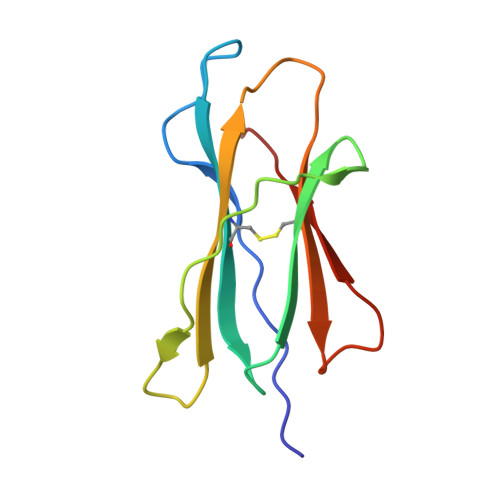
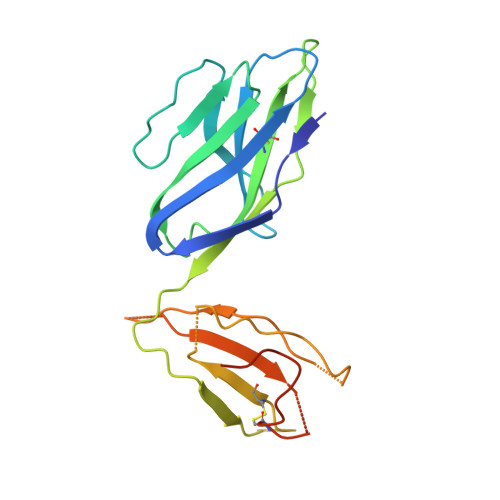
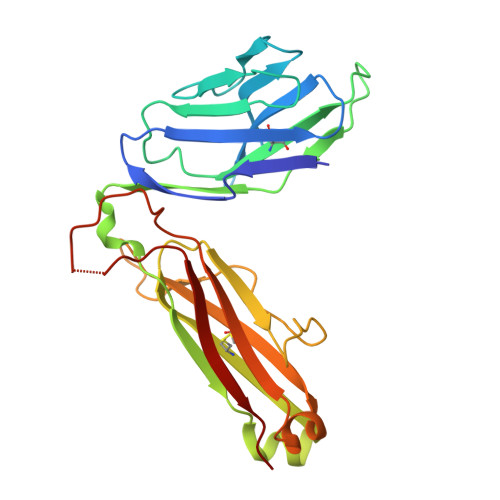
Mouse mucosal-associated invariant T cell receptor recognition of MR1 presenting the vitamin B metabolite, 5-(2-oxopropylideneamino)-6-d-ribitylaminouracil.
Ciacchi, L., Mak, J.Y.W., Le, J.P., Fairlie, D.P., McCluskey, J., Corbett, A.J., Rossjohn, J., Awad, W.(2024) J Biological Chem 300: 107229-107229
- PubMed: 38537698 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbc.2024.107229
- Primary Citation Related Structures:
8VZ8, 8VZ9 - PubMed Abstract:
Mucosal-associated invariant T (MAIT) cells can elicit immune responses against riboflavin-based antigens presented by the evolutionary conserved MHC class I related protein, MR1. While we have an understanding of the structural basis of human MAIT cell receptor (TCR) recognition of human MR1 presenting a variety of ligands, how the semi-invariant mouse MAIT TCR binds mouse MR1-ligand remains unknown. Here, we determine the crystal structures of 2 mouse TRAV1-TRBV13-2 + MAIT TCR-MR1-5-OP-RU ternary complexes, whose TCRs differ only in the composition of their CDR3β loops. These mouse MAIT TCRs mediate high affinity interactions with mouse MR1-5-OP-RU and cross-recognize human MR1-5-OP-RU. Similarly, a human MAIT TCR could bind mouse MR1-5-OP-RU with high affinity. This cross-species recognition indicates the evolutionary conserved nature of this MAIT TCR-MR1 axis. Comparing crystal structures of the mouse versus human MAIT TCR-MR1-5-OP-RU complexes provides structural insight into the conserved nature of this MAIT TCR-MR1 interaction and conserved specificity for the microbial antigens, whereby key germline-encoded interactions required for MAIT activation are maintained. This is an important consideration for the development of MAIT cell-based therapeutics that will rely on preclinical mouse models of disease.
- Infection and Immunity Program, Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria, Australia.
Organizational Affiliation: