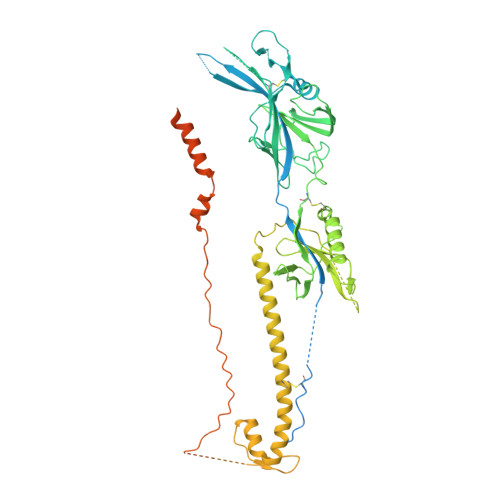
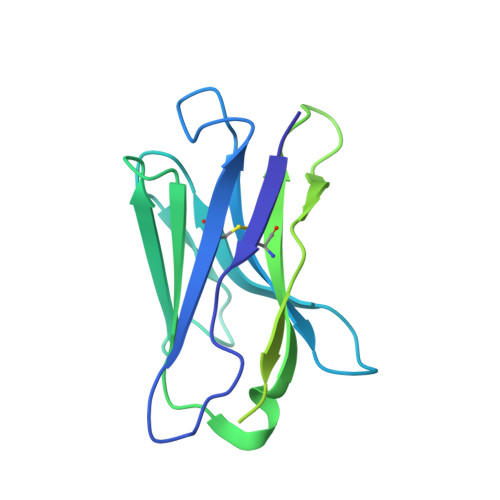
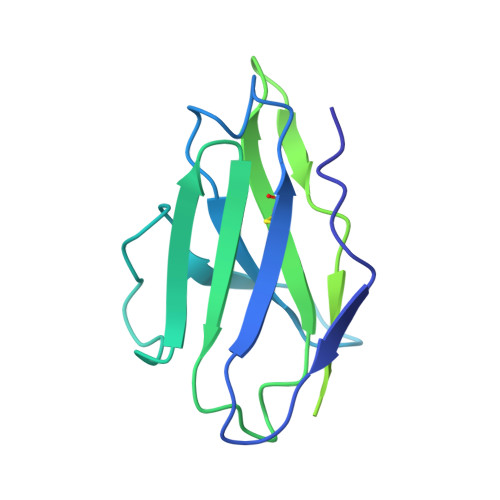
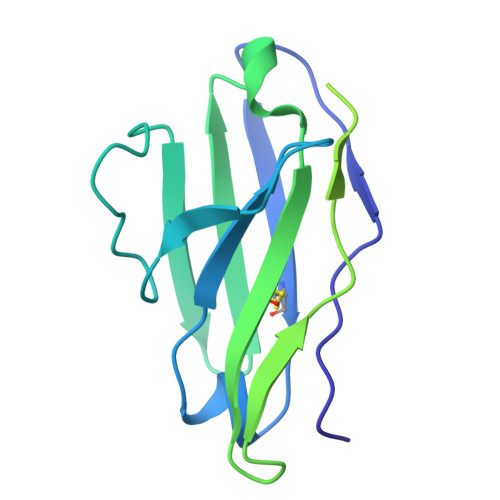
Structure-based design of a soluble human cytomegalovirus glycoprotein B antigen stabilized in a prefusion-like conformation.
Sponholtz, M.R., Byrne, P.O., Lee, A.G., Ramamohan, A.R., Goldsmith, J.A., McCool, R.S., Zhou, L., Johnson, N.V., Hsieh, C.L., Connors, M., Karthigeyan, K.P., Crooks, C.M., Fuller, A.S., Campbell, J.D., Permar, S.R., Maynard, J.A., Yu, D., Bottomley, M.J., McLellan, J.S.(2024) Proc Natl Acad Sci U S A 121: e2404250121-e2404250121
- PubMed: 39231203 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2404250121
- Primary Citation Related Structures:
8VYM, 8VYN - PubMed Abstract:
Human cytomegalovirus (HCMV) glycoprotein B (gB) is a class III membrane fusion protein required for viral entry. HCMV vaccine candidates containing gB have demonstrated moderate clinical efficacy, but no HCMV vaccine has been approved. Here, we used structure-based design to identify and characterize amino acid substitutions that stabilize gB in its metastable prefusion conformation. One variant containing two engineered interprotomer disulfide bonds and two cavity-filling substitutions (gB-C7), displayed increased expression and thermostability. A 2.8 Å resolution cryoelectron microscopy structure shows that gB-C7 adopts a prefusion-like conformation, revealing additional structural elements at the membrane-distal apex. Unlike previous observations for several class I viral fusion proteins, mice immunized with postfusion or prefusion-stabilized forms of soluble gB protein displayed similar neutralizing antibody titers, here specifically against an HCMV laboratory strain on fibroblasts. Collectively, these results identify initial strategies to stabilize class III viral fusion proteins and provide tools to probe gB-directed antibody responses.
- Department of Molecular Biosciences, The University of Texas at Austin, Austin, TX 78712.
Organizational Affiliation: