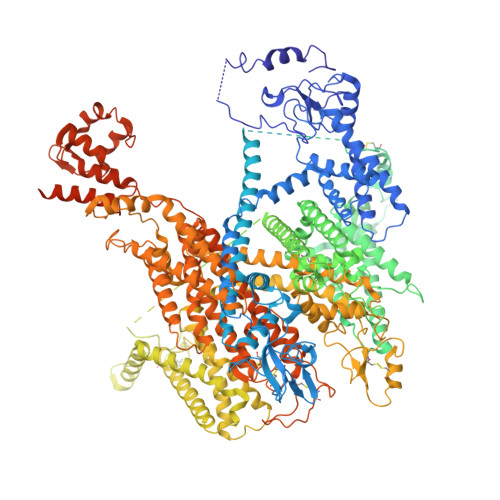
Structural basis of human Na v 1.5 gating mechanisms.
Biswas, R., Lopez-Serrano, A.L., Purohit, A., Ramirez-Navarro, A., Huang, H.L., Grandinetti, G., Cheng, X., Heissler, S.M., Deschenes, I., Chinthalapudi, K.(2025) Proc Natl Acad Sci U S A 122: e2416181122-e2416181122
- PubMed: 40366698 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2416181122
- Primary Citation Related Structures:
8VYJ, 8VYK - PubMed Abstract:
Voltage-gated Na v 1.5 channels are central to the generation and propagation of cardiac action potentials. Aberrations in their function are associated with a wide spectrum of cardiac diseases including arrhythmias and heart failure. Despite decades of progress in Na v 1.5 biology, the lack of structural insights into intracellular regions has hampered our understanding of its gating mechanisms. Here, we present two cryo-EM structures of human Na v 1.5 in open states, revealing sequential conformational changes in gating charges of the voltage-sensing domains (VSDs) and several intracellular regions. Despite the channel being in the open state, these structures show repositioning, but no dislodging of the IFM motif in the receptor site. Molecular dynamics analyses show our structures with CTD conduct Na + ions. Notably, our structural findings highlight a dynamic C-terminal domain (CTD) and III-IV linker interaction, which regulates the conformation of VSDs and pore opening. Electrophysiological studies confirm that disrupting this interaction alters fast inactivation of Na v 1.5. Together, our structure-function studies establish a foundation for understanding the gating mechanisms of Na v 1.5 and the mechanisms underlying CTD-related channelopathies.
- Department of Physiology and Cell Biology, Dorothy M. Davis Heart and Lung Research Institute, College of Medicine, The Ohio State University, Columbus, OH 43210.
Organizational Affiliation: