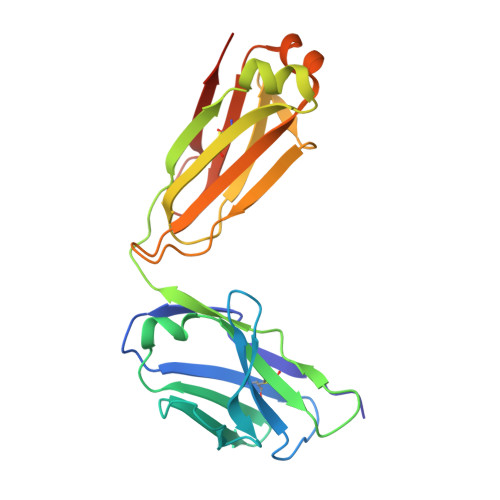
Structural, Biophysical, and Computational Studies of a Murine Light Chain Dimer.
Arriaza, R.H., Kapingidza, A.B., Dolamore, C., Khatri, K., O'Malley, A., Glesner, J., Wuenschmann, S., Hyduke, N.P., Easley, W., Chhiv, C., Pomes, A., Chruszcz, M.(2024) Molecules 29
- PubMed: 38930950
- DOI: https://doi.org/10.3390/molecules29122885
- Primary Citation of Related Structures:
8VY6 - PubMed Abstract:
Antibodies are widely used in medicinal and scientific research due to their ability to bind to a specific antigen. Most often, antibodies are composed of heavy and light chain domains. Under physiological conditions, light chains are produced in excess, as compared to the heavy chain. It is now known that light chains are not silent partners of the heavy chain and can modulate the immune response independently. In this work, the first crystal structure of a light chain dimer originating from mice is described. It represents the light chain dimer of 6A8, a monoclonal antibody specific to the allergen Der f 1. Building on the unexpected occurrence of this kind of dimer, we have demonstrated that this light chain is stable in solution alone. Moreover, enzyme-linked immunosorbent assays (ELISA) have revealed that, when the light chain is not partnered to its corresponding heavy chain, it interacts non-specifically with a wide range of proteins. Computational studies were used to provide insight on the role of the 6A8 heavy chain domain in the specific binding to Der f 1. Overall, this work demonstrates and supports the ongoing notion that light chains can function by themselves and are not silent partners of heavy chains.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Michigan State University, East Lansing, MI 48864, USA.