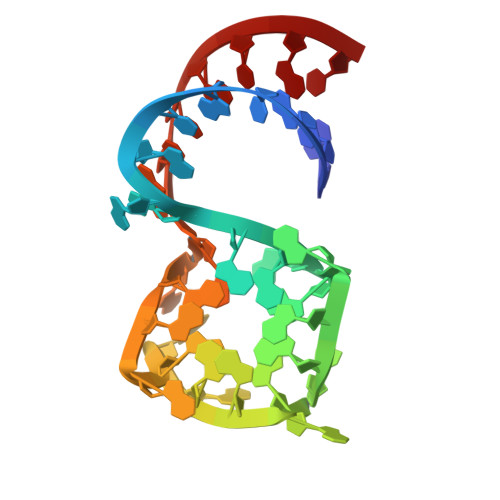
Structure-informed design of an ultrabright RNA-activated fluorophore.
Yang, M., Prestwood, P.R., Passalacqua, L.F.M., Balaratnam, S., Fullenkamp, C.R., Arney, J.W., Weeks, K.M., Ferre-D'Amare, A., Schneekloth Jr., J.S.(2025) Nat Chem 17: 1188-1195
- PubMed: 40437193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41557-025-01832-w
- Primary Citation Related Structures:
8VXX, 8VXZ, 8VY0, 8VY1 - PubMed Abstract:
RNA-based fluorogenic aptamers, such as Mango, are uniquely powerful tools for imaging RNA that activate the fluorescence of a weakly or non-fluorescent small molecule when bound. A central challenge has been to develop brighter, more specific and high-affinity aptamer-ligand systems for cellular imaging. Here we report an ultrabright fluorophore for the Mango II system discovered using a structure-informed, fragment-based small-molecule microarray approach. This dye-termed SALAD1 (structure-informed, array-enabled LigAnD 1)-exhibits subnanomolar aptamer affinity and 3.5-fold brighter fluorescence than Mango II-TO1-biotin pair, a widely used fluorogenic system. Performance was improved by modulating RNA-dye molecular recognition without altering the fluorophore's π-system. High-resolution X-ray structures reveal the binding mode for SALAD1, which exhibits improved pocket occupancy, a more defined binding pose and a unique bonding interaction with potassium. SALAD1 is cell-permeable and facilitates improved in-cell confocal RNA imaging. This work introduces an additional RNA-activated fluorophore demonstrating how fragment-based ligand discovery can be used to create high-performance ligands for RNA targets.
- Chemical Biology Laboratory, Center for Cancer Research, National Cancer Institute, Frederick, MD, USA.
Organizational Affiliation: