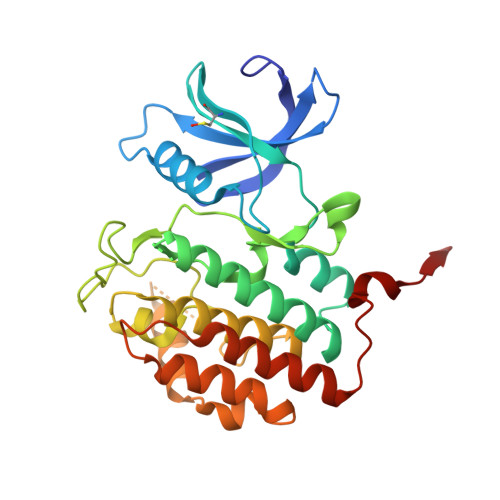
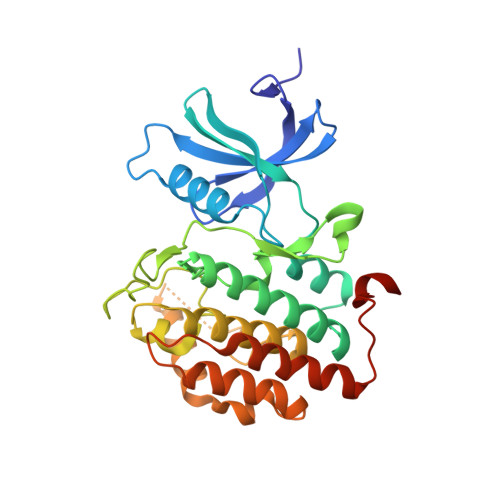
Structure-Based Optimization of Selective and Brain Penetrant CK1 delta Inhibitors for the Treatment of Circadian Disruptions.
McCarver, S., Hanna, L., Samant, A., Thompson, A.A., Seierstad, M., Saha, A., Wu, D., Lord, B., Sutton, S.W., Shah, V., Milligan, C.M., Wennerholm, M., Shelton, J., Lebold, T.P., Shireman, B.T.(2024) ACS Med Chem Lett 15: 486-492
- PubMed: 38628796 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.3c00523
- Primary Citation Related Structures:
8VXD, 8VXE, 8VXF - PubMed Abstract:
Neuropsychiatric disorders such as major depressive disorders and schizophrenia are often associated with disruptions to the normal 24 h sleep wake cycle. Casein kinase 1 (CK1δ) is an integral part of the molecular machinery that regulates circadian rhythms. Starting from a cluster of bicyclic pyrazoles identified from a virtual screening effort, we utilized structure-based drug design to identify and reinforce a unique "hinge-flip" binding mode that provides a high degree of selectivity for CK1δ versus the kinome. Pharmacokinetics, brain exposure, and target engagement as measured by ex vivo autoradiography are described for advanced analogs.
- Janssen Research and Development, San Diego, California 92121-1126, United States.
Organizational Affiliation: