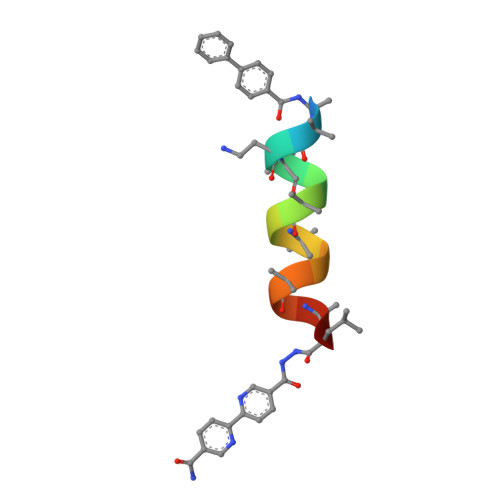
Diverse Proteomimetic Frameworks via Rational Design of pi-Stacking Peptide Tectons.
Ganatra, P., Wang, D.F., Ganatra, V., Dang, V.T., Nguyen, A.I.(2024) J Am Chem Soc 146: 22236-22246
- PubMed: 39096501 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.4c03094
- Primary Citation Related Structures:
8VOG, 8VP7, 8VPC, 8VPD, 8VPE, 8VPJ, 8VPS, 8VPT, 8VPX, 8VPY, 8VPZ, 8VQ0, 8VSF, 8VT8, 8VW7, 8VW8, 8VXS - PubMed Abstract:
Peptide-based frameworks aim to integrate protein architecture into solid-state materials using simpler building blocks. Despite the growing number of peptide frameworks, there are few strategies to rationally engineer essential properties like pore size and shape. Designing peptide assemblies is generally hindered by the difficulty of predicting complex networks of weak intermolecular interactions. Peptides conjugated to polyaromatic groups are a unique case where assembly appears to be strongly driven by π-π interactions, suggesting that rationally adjusting the geometry of the π-stackers could create novel structures. Here, we report peptide elongation as a simple mechanism to predictably tune the angle between the π-stacking groups to produce a remarkable diversity of pore shapes and sizes, including some that are mesoporous. Notably, rapid jumps in pore size and shape can occur with just a single amino acid insertion. The geometry of the π-stacking residues also significantly influences framework structure, representing an additional dimension for tuning. Lastly, sequence identity can also indirectly modulate the π-π interactions. By correlating each of these factors with detailed crystallographic data, we find that, despite the complexity of peptide structure, the shape and polarity of the tectons are straightforward predictors of framework structure. These guidelines are expected to accelerate the development of advanced porous materials with protein-like capabilities.
- Department of Chemistry, University of Illinois Chicago, Chicago, Illinois 60607, United States.
Organizational Affiliation: