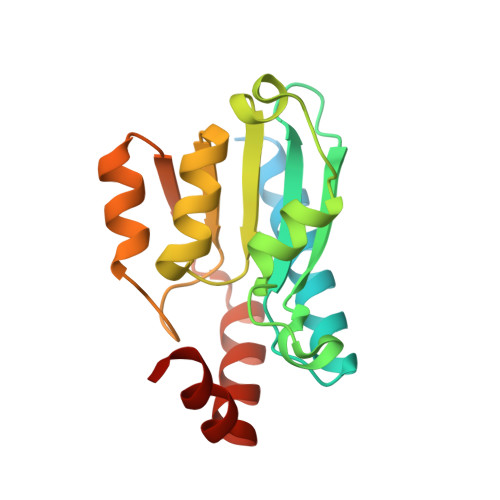
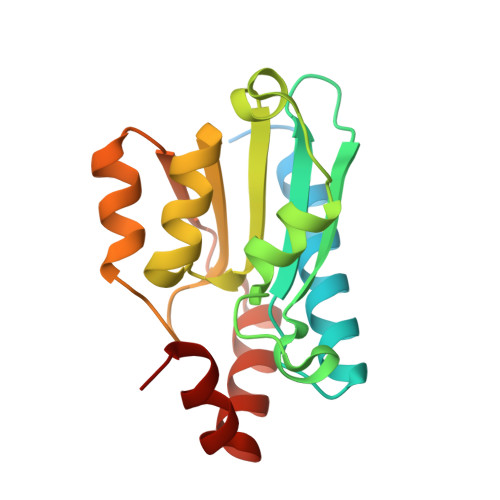
Compensatory mutations potentiate constructive neutral evolution by gene duplication.
Despres, P.C., Dube, A.K., Picard, M.E., Grenier, J., Shi, R., Landry, C.R.(2024) Science 385: 770-775
- PubMed: 39146405 Search on PubMed
- DOI: https://doi.org/10.1126/science.ado5719
- Primary Citation Related Structures:
8VLJ, 8VLK, 8VLL, 8VLM - PubMed Abstract:
The functions of proteins generally depend on their assembly into complexes. During evolution, some complexes have transitioned from homomers encoded by a single gene to heteromers encoded by duplicate genes. This transition could occur without adaptive evolution through intermolecular compensatory mutations. Here, we experimentally duplicated and evolved a homodimeric enzyme to determine whether and how this could happen. We identified hundreds of deleterious mutations that inactivate individual homodimers but produce functional enzymes when coexpressed as duplicated proteins that heterodimerize. The structure of one such heteromer reveals how both losses of function are buffered through the introduction of asymmetry in the complex that allows them to subfunctionalize. Constructive neutral evolution can thus occur by gene duplication followed by only one deleterious mutation per duplicate.
- Département de Biochimie, de Microbiologie et de Bio-informatique, Faculté des Sciences et de Génie Université Laval, Québec, QC G1V 0A6, Canada.
Organizational Affiliation: