Exploring Long Arm Amide-Linked Side Chains in the Design of Antifungal Azole Inhibitors of Sterol 14 alpha-Demethylase (CYP51).
Alsulaimany, M., Keniya, M.V., Alanazi, R.S., N Ruma, Y., Hughes, C.S., Jones, A.T., Tyndall, J.D.A., Parker, J.E., Monk, B.C., Simons, C.(2025) J Med Chem 68: 10781-10799
- PubMed: 40403151 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.4c02922
- Primary Citation Related Structures:
8VK6 - PubMed Abstract:
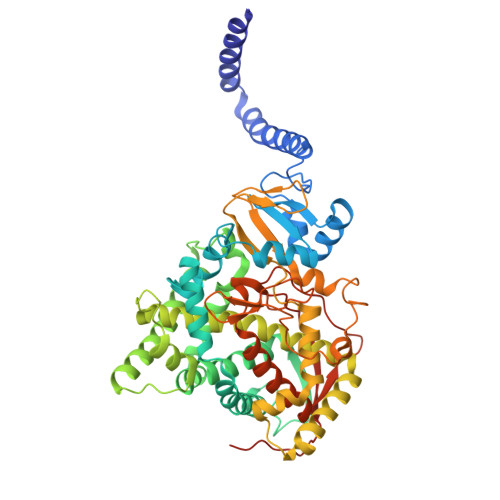
The rise in fungal drug resistance has exacerbated the treatment of invasive fungal infections, most commonly caused by Candida . This research describes the synthesis of extended "long-arm" azole antifungals that were evaluated against wild-type and resistant fungal species. Biphenyl derivative 22 was the most effective derivative, displaying potent inhibitory activity against Saccharomyces , Candida , and Cryptococcus CYP51 enzymes, including in resistant strains, in comparison with posaconazole. The X-ray crystal structure of S - 22 complexed with S. cerevisiae CYP51 showed a hydrogen bond between the oxygen of the trifluoromethoxy group of S- 22 and the His381 side chain of S. cerevisiae CYP51, which is postulated to contribute significantly to its binding, and stabilization in the presence of the S. cerevisiae CYP51 Y140F/H, C. parapsilosis and C. auris CYP51 Y132F mutations and the C. auris K143R mutation. Computational studies and IC 50 evaluation of compound 22 vs C. albicans wild-type, Y132F, and Y132H/K143 mutant strains supported MIC observations.
- School of Pharmacy and Pharmaceutical Sciences, Cardiff University, King Edward VII Avenue, Cardiff CF10 3NB, U.K.
Organizational Affiliation: