Defining bottlenecks and opportunities for Lassa virus neutralization by structural profiling of vaccine-induced polyclonal antibody responses.
Brouwer, P.J.M., Perrett, H.R., Beaumont, T., Nijhuis, H., Kruijer, S., Burger, J.A., Bontjer, I., Lee, W.H., Ferguson, J.A., Schauflinger, M., Muller-Krauter, H., Sanders, R.W., Strecker, T., van Gils, M.J., Ward, A.B.(2024) Cell Rep 43: 114708-114708
- PubMed: 39243373 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2024.114708
- Primary Citation Related Structures:
8TYC, 8TYE, 8VCV, 8VE8, 9CJ7, 9CJ8, 9CK7, 9CK8 - PubMed Abstract:
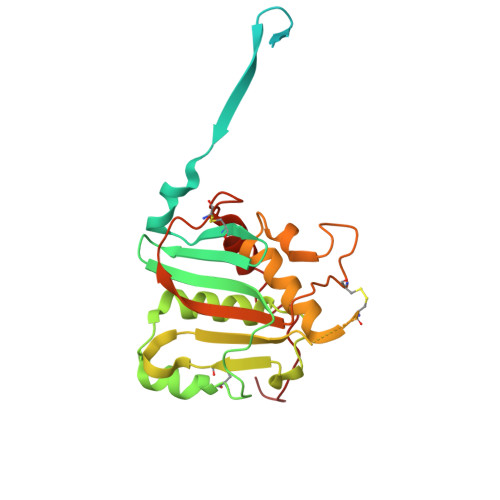
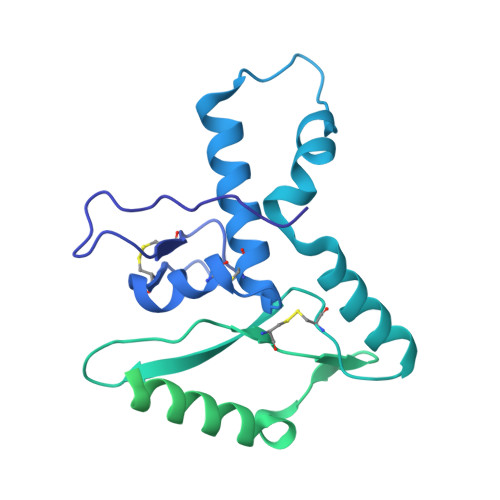
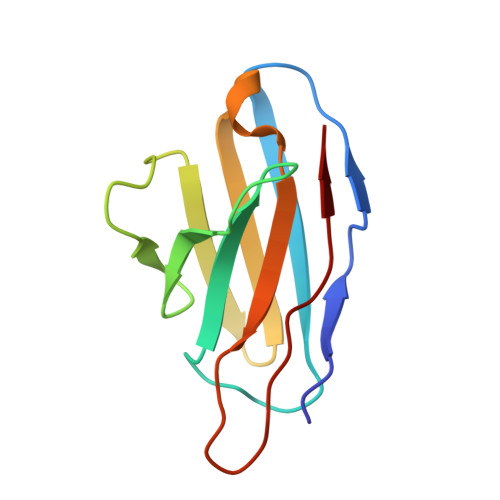
Lassa fever continues to be a major public health burden in West Africa, yet effective therapies or vaccines are lacking. The isolation of protective neutralizing antibodies against the Lassa virus glycoprotein complex (GPC) justifies the development of vaccines that can elicit strong neutralizing antibody responses. However, Lassa vaccine candidates have generally been unsuccessful at doing so, and the associated antibody responses to these vaccines remain poorly characterized. Here, we establish an electron microscopy-based epitope mapping workflow that enables high-resolution structural characterization of polyclonal antibodies to the GPC. By applying this method to rabbits vaccinated with a recombinant GPC vaccine and a GPC-derived virus-like particle, we reveal determinants of neutralization that involve epitopes of the GPC-A competition cluster. Furthermore, by identifying undescribed immunogenic off-target epitopes, we expose the challenges that recombinant GPC vaccines face. By enabling detailed polyclonal antibody characterization, our work ushers in a next generation of more rational Lassa vaccine design.
- Department of Integrative Structural and Computational Biology, Scripps Research, La Jolla, CA 92037, USA.
Organizational Affiliation: