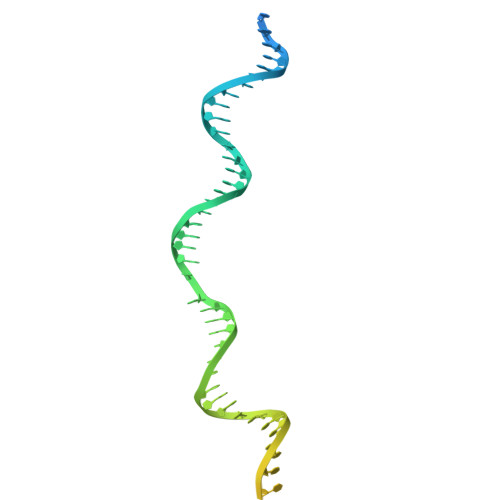
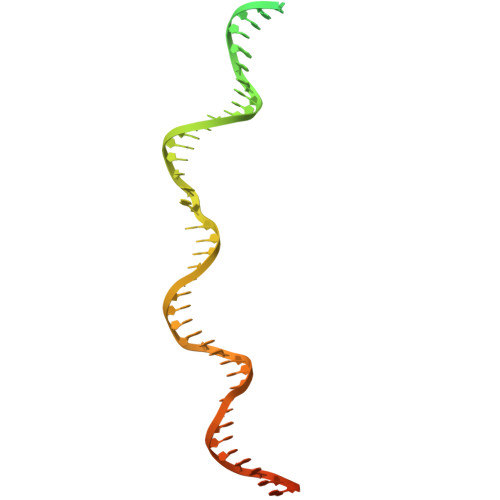
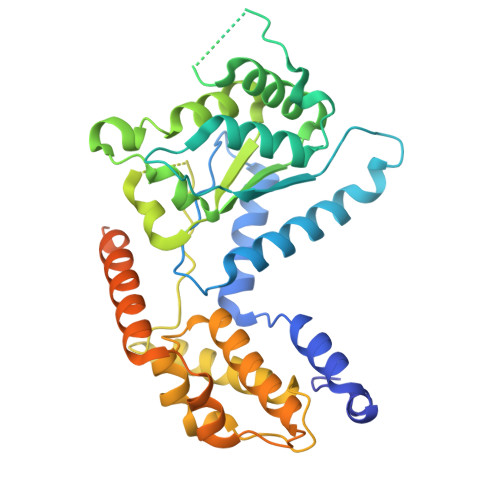
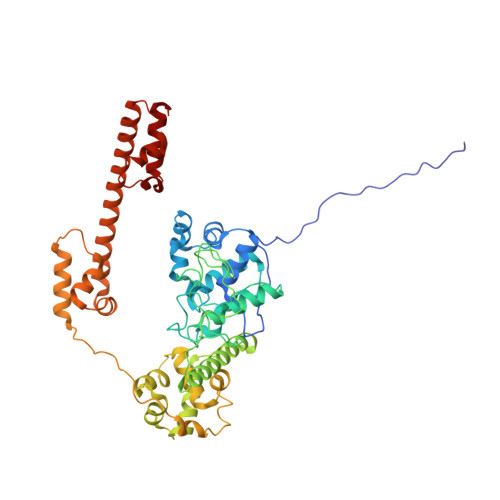
Structure of TnsABCD transpososome reveals mechanisms of targeted DNA transposition.
Wang, S., Siddique, R., Hall, M.C., Rice, P.A., Chang, L.(2024) Cell 187: 6865-6881.e16
- PubMed: 39383864 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2024.09.023
- Primary Citation Related Structures:
8V32, 9BW1 - PubMed Abstract:
Tn7-like transposons are characterized by their ability to insert specifically into host chromosomes. Recognition of the attachment (att) site by TnsD recruits the TnsABC proteins to form the transpososome and facilitate transposition. Although this pathway is well established, atomic-level structural insights of this process remain largely elusive. Here, we present the cryo-electron microscopy (cryo-EM) structures of the TnsC-TnsD-att DNA complex and the TnsABCD transpososome from the Tn7-like transposon in Peltigera membranacea cyanobiont 210A, a type I-B CRISPR-associated transposon. Our structures reveal a striking bending of the att DNA, featured by the intercalation of an arginine side chain of TnsD into a CC/GG dinucleotide step. The TnsABCD transpososome structure reveals TnsA-TnsB interactions and demonstrates that TnsC not only recruits TnsAB but also directly participates in the transpososome assembly. These findings provide mechanistic insights into targeted DNA insertion by Tn7-like transposons, with implications for improving the precision and efficiency of their genome-editing applications.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA.
Organizational Affiliation: