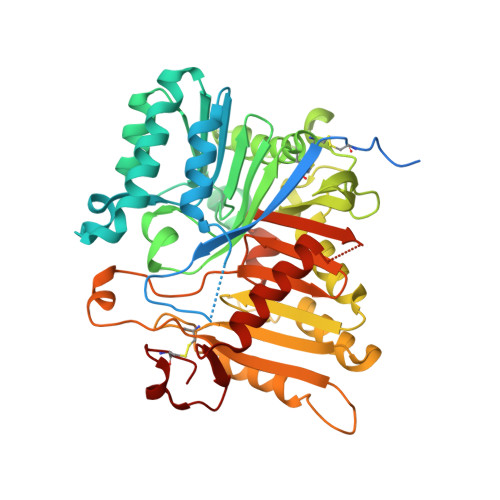
Structural and mechanistic insights into disease-associated endolysosomal exonucleases PLD3 and PLD4.
Yuan, M., Peng, L., Huang, D., Gavin, A., Luan, F., Tran, J., Feng, Z., Zhu, X., Matteson, J., Wilson, I.A., Nemazee, D.(2024) Structure 32: 766-779.e7
- PubMed: 38537643 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2024.02.019
- Primary Citation Related Structures:
8V05, 8V06, 8V07, 8V08 - PubMed Abstract:
Endolysosomal exonucleases PLD3 and PLD4 (phospholipases D3 and D4) are associated with autoinflammatory and autoimmune diseases. We report structures of these enzymes, and the molecular basis of their catalysis. The structures reveal an intra-chain dimer topology forming a basic active site at the interface. Like other PLD superfamily members, PLD3 and PLD4 carry HxKxxxxD/E motifs and participate in phosphodiester-bond cleavage. The enzymes digest ssDNA and ssRNA in a 5'-to-3' manner and are blocked by 5'-phosphorylation. We captured structures in apo, intermediate, and product states and revealed a "link-and-release" two-step catalysis. We also unexpectedly demonstrated phosphatase activity via a covalent 3-phosphohistidine intermediate. PLD4 contains an extra hydrophobic clamp that stabilizes substrate and could affect oligonucleotide substrate preference and product release. Biochemical and structural analysis of disease-associated mutants of PLD3/4 demonstrated reduced enzyme activity or thermostability and the possible basis for disease association. Furthermore, these findings provide insight into therapeutic design.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA. Electronic address: myuan@scripps.edu.
Organizational Affiliation: