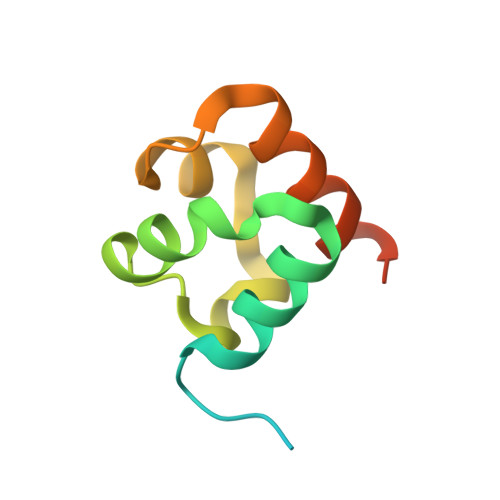
Structural analysis of the SAM domain of the Arabidopsis mitochondrial tRNA import receptor.
Olasz, B., Smithers, L., Evans, G.L., Anandan, A., Murcha, M.W., Vrielink, A.(2024) J Biological Chem 300: 107258-107258
- PubMed: 38582448 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbc.2024.107258
- Primary Citation Related Structures:
8UCY, 8UCZ, 8UD0 - PubMed Abstract:
Mitochondria are membrane-bound organelles of endosymbiotic origin with limited protein-coding capacity. The import of nuclear-encoded proteins and nucleic acids is required and essential for maintaining organelle mass, number, and activity. As plant mitochondria do not encode all the necessary tRNA types required, the import of cytosolic tRNA is vital for organelle maintenance. Recently, two mitochondrial outer membrane proteins, named Tric1 and Tric2, for tRNA import component, were shown to be involved in the import of cytosolic tRNA. Tric1/2 binds tRNA ala via conserved residues in the C-terminal Sterile Alpha Motif (SAM) domain. Here we report the X-ray crystal structure of the Tric1 SAM domain. We identified the ability of the SAM domain to form a helical superstructure with six monomers per helical turn and key amino acid residues responsible for its formation. We determined that the oligomerization of the Tric1 SAM domain may play a role in protein function whereby mutation of Gly241 introducing a larger side chain at this position disrupted the oligomer and resulted in the loss of RNA binding capability. Furthermore, complementation of Arabidopsis thaliana Tric1/2 knockout lines with a mutated Tric1 failed to restore the defective plant phenotype. AlphaFold2 structure prediction of both the SAM domain and Tric1 support a cyclic pentameric or hexameric structure. In the case of a hexameric structure, a pore of sufficient dimensions to transfer tRNA across the mitochondrial membrane is observed. Our results highlight the importance of oligomerization of Tric1 for protein function.
- School of Molecular Sciences, The University of Western Australia, Perth, Western Australia, Australia.
Organizational Affiliation: