Structure of isomerase domain of the nan-regulatory protein (NanR) from Streptococcus pneumoniae
Wood, D.M., Dobson, R.C.J.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| MurR/RpiR family transcriptional regulator | A, B [auth C], C [auth D], D [auth B] | 283 | Streptococcus pneumoniae | Mutation(s): 0 | 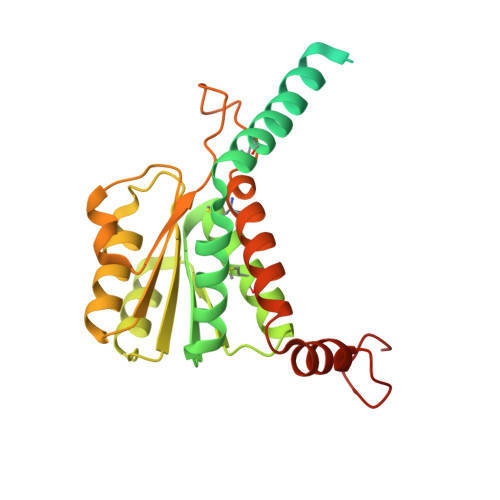 |
UniProt | |||||
Find proteins for A0A064C3N8 (Streptococcus pneumoniae) Explore A0A064C3N8 Go to UniProtKB: A0A064C3N8 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A064C3N8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| BMX (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth A], G [auth C], I [auth D], K [auth B] | 2-acetamido-2-deoxy-6-O-phosphono-alpha-D-mannopyranose C8 H16 N O9 P BRGMHAYQAZFZDJ-UOLFYFMNSA-N |  | ||
| NA Download:Ideal Coordinates CCD File | F [auth A], H [auth C], J [auth D], L [auth B] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A, B [auth C], C [auth D], D [auth B] | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 147.577 | α = 90 |
| b = 147.577 | β = 90 |
| c = 82.311 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| PHENIX | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Royal Society of New Zealand | New Zealand | UOC1506 |
| Ministry of Business, Innovation and Employment (New Zealand) | New Zealand | UOCX1706 |