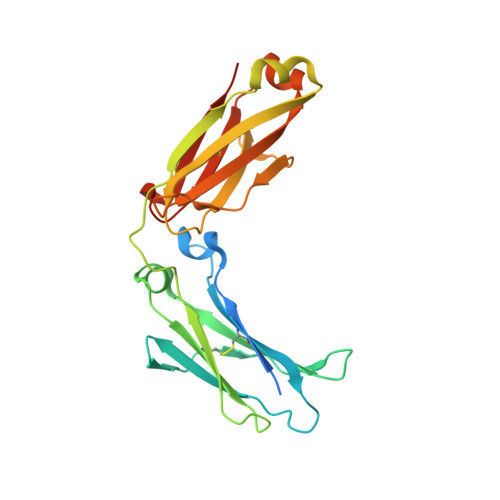
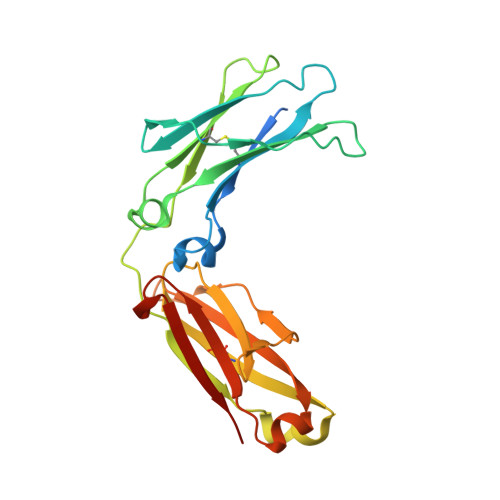
Combinatorially restricted computational design of protein-protein interfaces to produce IgG heterodimers.
Azzam, T., Du, J.J., Flowers, M.W., Ali, A.V., Hunn, J.C., Vijayvargiya, N., Knagaram, R., Bogacz, M., Maravillas, K.E., Sastre, D.E., Fields, J.K., Mirzaei, A., Pierce, B.G., Sundberg, E.J.(2024) Sci Adv 10: eadk8157-eadk8157
- PubMed: 38598628 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adk8157
- Primary Citation Related Structures:
8TTM, 8TUD - PubMed Abstract:
Redesigning protein-protein interfaces is an important tool for developing therapeutic strategies. Interfaces can be redesigned by in silico screening, which allows for efficient sampling of a large protein space before experimental validation. However, computational costs limit the number of combinations that can be reasonably sampled. Here, we present combinatorial tyrosine (Y)/serine (S) selection (combYSelect), a computational approach combining in silico determination of the change in binding free energy (ΔΔ G ) of an interface with a highly restricted library composed of just two amino acids, tyrosine and serine. We used combYSelect to design two immunoglobulin G (IgG) heterodimers-combYSelect1 (L368S/D399Y-K409S/T411Y) and combYSelect2 (D399Y/K447S-K409S/T411Y)-that exhibit near-optimal heterodimerization, without affecting IgG stability or function. We solved the crystal structures of these heterodimers and found that dynamic π-stacking interactions and polar contacts drive preferential heterodimeric interactions. Finally, we demonstrated the utility of our combYSelect heterodimers by engineering both a bispecific antibody and a cytokine trap for two unique therapeutic applications.
- Department of Biochemistry, Emory University School of Medicine, Atlanta, GA 30322, USA.
Organizational Affiliation: