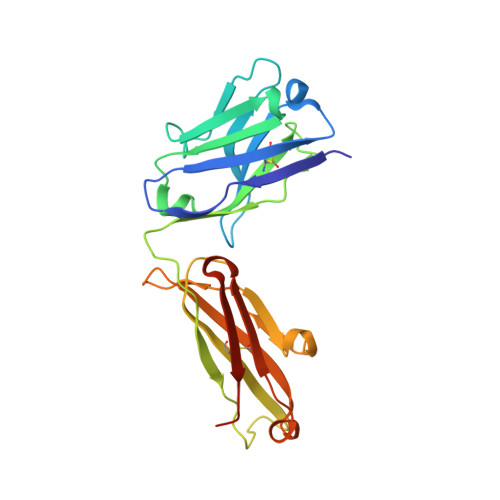
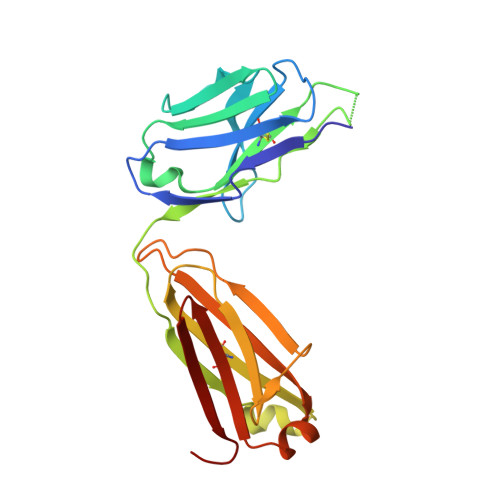
Engineered antigen-binding fragments for enhanced crystallization of antibody:antigen complexes.
Bruce, H.A., Singer, A.U., Filippova, E.V., Blazer, L.L., Adams, J.J., Enderle, L., Ben-David, M., Radley, E.H., Mao, D.Y.L., Pau, V., Orlicky, S., Sicheri, F., Kurinov, I., Atwell, S., Kossiakoff, A.A., Sidhu, S.S.(2024) Protein Sci 33: e4824-e4824
- PubMed: 37945533 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4824
- Primary Citation Related Structures:
8T58, 8T6I, 8T7F, 8T7G, 8T7I, 8T8I, 8T9Y, 8TRS, 8TRT, 8TS5 - PubMed Abstract:
The atomic-resolution structural information that X-ray crystallography can provide on the binding interface between a Fab and its cognate antigen is highly valuable for understanding the mechanism of interaction. However, many Fab:antigen complexes are recalcitrant to crystallization, making the endeavor a considerable effort with no guarantee of success. Consequently, there have been significant steps taken to increase the likelihood of Fab:antigen complex crystallization by altering the Fab framework. In this investigation, we applied the surface entropy reduction strategy coupled with phage-display technology to identify a set of surface substitutions that improve the propensity of a human Fab framework to crystallize. In addition, we showed that combining these surface substitutions with previously reported Crystal Kappa and elbow substitutions results in an extraordinary improvement in Fab and Fab:antigen complex crystallizability, revealing a strong synergistic relationship between these sets of substitutions. Through comprehensive Fab and Fab:antigen complex crystallization screenings followed by structure determination and analysis, we defined the roles that each of these substitutions play in facilitating crystallization and how they complement each other in the process.
- School of Pharmacy, University of Waterloo, Waterloo, Canada.
Organizational Affiliation: