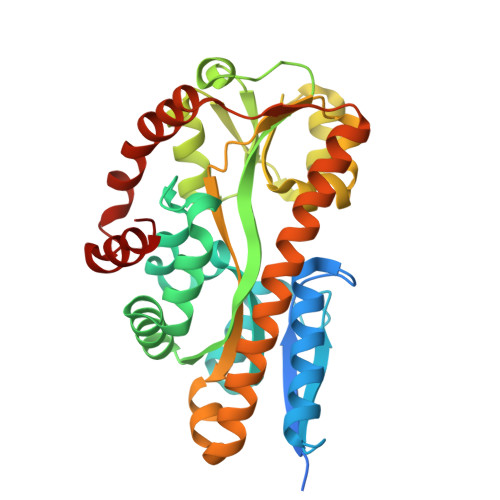
On the function of TRAP substrate-binding proteins: the isethionate-specific binding protein IseP.
Newton-Vesty, M.C., Currie, M.J., Davies, J.S., Panjikar, S., Sethi, A., Whitten, A.E., Tillett, Z.D., Wood, D.M., Wright, J.D., Love, M.J., Allison, T.M., Jamieson, S.A., Mace, P.D., North, R.A., Dobson, R.C.J.(2024) Biochem J 481: 1901-1920
- PubMed: 39560287 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BCJ20240540
- Primary Citation Related Structures:
8T9T, 8TE9, 8TQN, 8TRP - PubMed Abstract:
Bacteria evolve mechanisms to compete for limited resources and survive in new niches. Here we study the mechanism of isethionate import from the sulfate-reducing bacterium Oleidesulfovibrio alaskensis. The catabolism of isethionate by Desulfovibrio species has been implicated in human disease, due to hydrogen sulfide production, and has potential for industrial applications. O. alaskensis employs a tripartite ATP-independent periplasmic (TRAP) transporter (OaIsePQM) to import isethionate, which relies on the substrate-binding protein (OaIseP) to scavenge isethionate and deliver it to the membrane transporter component (OaIseQM) for import into the cell. We determined the binding affinity of isethionate to OaIseP by isothermal titration calorimetry, KD = 0.95 µM (68% CI = 0.6-1.4 µM), which is weaker compared with other TRAP substrate-binding proteins. The X-ray crystal structures of OaIseP in the ligand-free and isethionate-bound forms were obtained and showed that in the presence of isethionate, OaIseP adopts a closed conformation whereby two domains of the protein fold over the substrate. We serendipitously discovered two crystal forms with sulfonate-containing buffers (HEPES and MES) bound in the isethionate-binding site. However, these do not evoke domain closure, presumably because of the larger ligand size. Together, our data elucidate the molecular details of how a TRAP substrate-binding protein binds a sulfonate-containing substrate, rather than a typical carboxylate-containing substrate. These results may inform future antibiotic development to target TRAP transporters and provide insights into protein engineering of TRAP transporter substrate-binding proteins.
- Biomolecular Interaction Centre, School of Biological Sciences, MacDiarmid Institute for Advanced Materials and Nanotechnology, University of Canterbury, Christchurch 8140, New Zealand.
Organizational Affiliation: