Functional and structural basis of human parainfluenza virus type 3 neutralization with human monoclonal antibodies.
Suryadevara, N., Otrelo-Cardoso, A.R., Kose, N., Hu, Y.X., Binshtein, E., Wolters, R.M., Greninger, A.L., Handal, L.S., Carnahan, R.H., Moscona, A., Jardetzky, T.S., Crowe Jr., J.E.(2024) Nat Microbiol 9: 2128-2143
- PubMed: 38858594 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-024-01722-w
- Primary Citation Related Structures:
8TQI, 8TQK - PubMed Abstract:
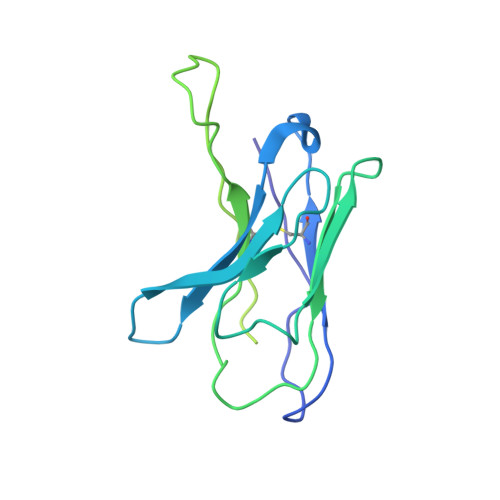
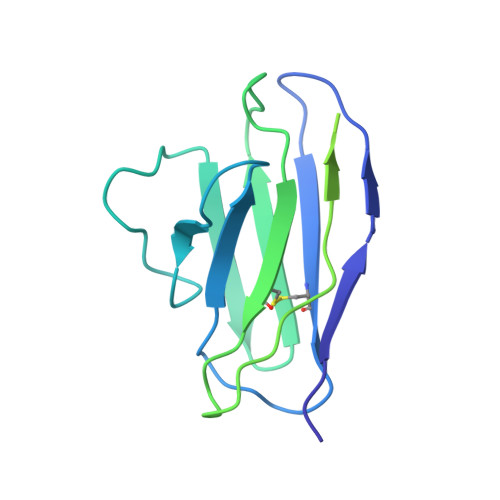
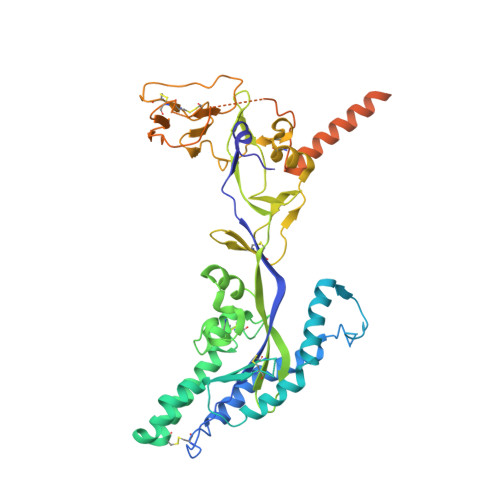
Human parainfluenza virus type 3 (hPIV3) is a respiratory pathogen that can cause severe disease in older people and infants. Currently, vaccines against hPIV3 are in clinical trials but none have been approved yet. The haemagglutinin-neuraminidase (HN) and fusion (F) surface glycoproteins of hPIV3 are major antigenic determinants. Here we describe naturally occurring potently neutralizing human antibodies directed against both surface glycoproteins of hPIV3. We isolated seven neutralizing HN-reactive antibodies and a pre-fusion conformation F-reactive antibody from human memory B cells. One HN-binding monoclonal antibody (mAb), designated PIV3-23, exhibited functional attributes including haemagglutination and neuraminidase inhibition. We also delineated the structural basis of neutralization for two HN and one F mAbs. MAbs that neutralized hPIV3 in vitro protected against infection and disease in vivo in a cotton rat model of hPIV3 infection, suggesting correlates of protection for hPIV3 and the potential clinical utility of these mAbs.
- Vanderbilt Vaccine Center, Vanderbilt University Medical Center, Nashville, TN, USA.
Organizational Affiliation: