Crystal structure of Fab M142 in complex with MHC-I (H2-Dd)
Jiang, J., Natarajan, K., Boyd, L.F., Margulies, D.H.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| H2 class I histocompatibility antigen D-d alpha chain (H2-Dd) | 273 | Mus musculus | Mutation(s): 0 Gene Names: H2-D1 | 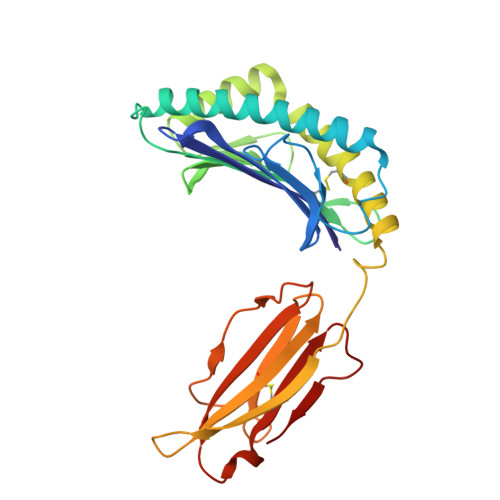 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P01900 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-2-microglobulin | 98 | Mus musculus | Mutation(s): 0 Gene Names: B2m | 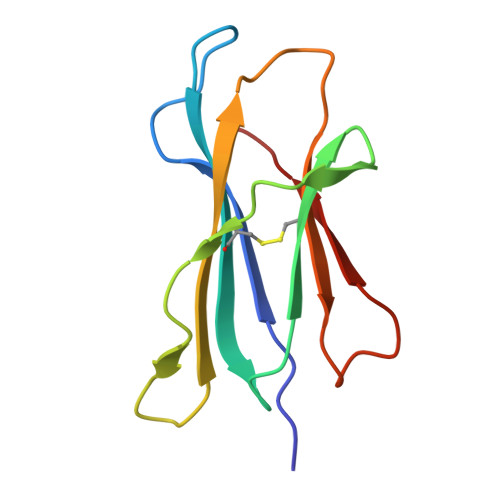 | |
UniProt & NIH Common Fund Data Resources | |||||
IMPC: MGI:88127 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P01887 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab M142 Heavy Chain | 222 | Rattus norvegicus | Mutation(s): 0 | 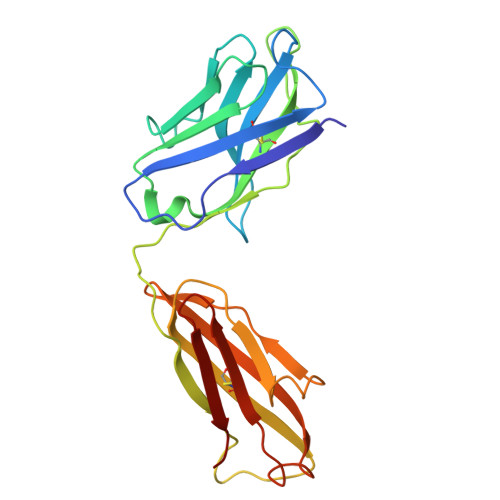 | |
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab M142 Light Chain | D, I [auth L] | 213 | Rattus norvegicus | Mutation(s): 0 | 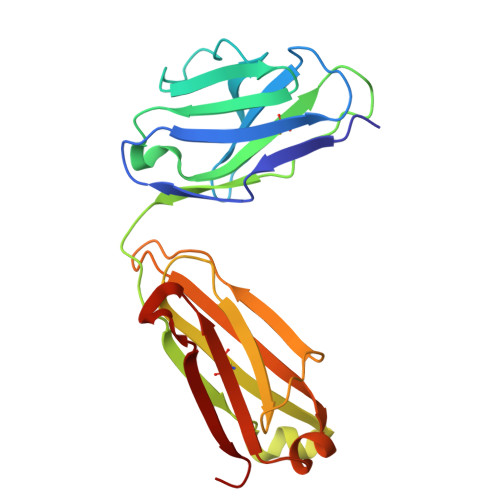 |
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| HV1: HIV-1 P18-I10 | G, J [auth P] | 10 | synthetic construct | Mutation(s): 0 |  |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 121.552 | α = 90 |
| b = 121.552 | β = 90 |
| c = 327.199 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | -- |