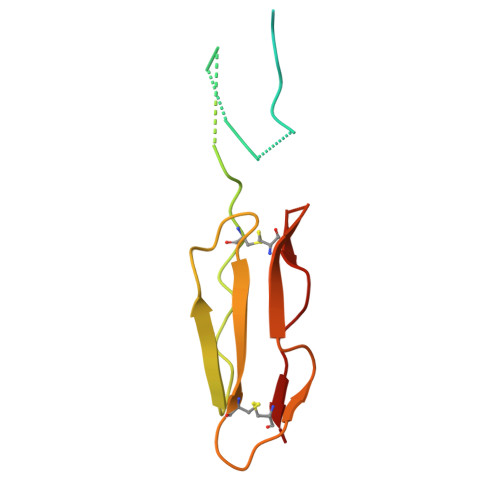
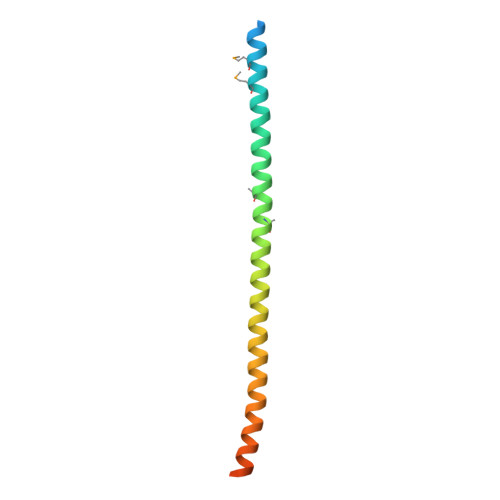Conservation of C4BP-binding sequence patterns in Streptococcus pyogenes M and Enn proteins.
Kolesinski, P., McGowan, M., Botteaux, A., Smeesters, P.R., Ghosh, P.(2024) J Biological Chem 300: 107478-107478
- PubMed: 38879009 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2024.107478
- Primary Citation Related Structures:
8TCB, 8TGT - PubMed Abstract:
Antigenically sequence variable M proteins of the major bacterial pathogen Streptococcus pyogenes (Strep A) are responsible for recruiting human C4b-binding protein (C4BP) to the bacterial surface, which enables Strep A to evade destruction by the immune system. The most sequence divergent portion of M proteins, the hypervariable region (HVR), is responsible for binding C4BP. Structural evidence points to the conservation of two C4BP-binding sequence patterns (M2 and M22) in the HVR of numerous M proteins, with this conservation applicable to vaccine immunogen design. These two patterns, however, only partially explain C4BP binding by Strep A. Here, we identified several M proteins that lack these patterns but still bind C4BP and determined the structures of two, M68 and M87 HVRs, in complex with a C4BP fragment. Mutagenesis of these M proteins led to the identification of amino acids that are crucial for C4BP binding, enabling formulation of new C4BP-binding patterns. Mutagenesis was also carried out on M2 and M22 proteins to refine or generate experimentally grounded C4BP-binding patterns. The M22 pattern was the most prevalent among M proteins, followed by the M87 and M2 patterns, while the M68 pattern was rare. These patterns, except for M68, were also evident in numerous M-like Enn proteins. Binding of C4BP via these patterns to Enn proteins was verified. We conclude that C4BP-binding patterns occur frequently in Strep A strains of differing M types, being present in their M or Enn proteins, or frequently both, providing further impetus for their use as vaccine immunogens.
- Department of Chemistry and Biochemistry, University of California, San Diego La Jolla, California, USA.
Organizational Affiliation: