Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines.
Del Moral-Sanchez, I., Wee, E.G., Xian, Y., Lee, W.H., Allen, J.D., Torrents de la Pena, A., Froes Rocha, R., Ferguson, J., Leon, A.N., Koekkoek, S., Schermer, E.E., Burger, J.A., Kumar, S., Zwolsman, R., Brinkkemper, M., Aartse, A., Eggink, D., Han, J., Yuan, M., Crispin, M., Ozorowski, G., Ward, A.B., Wilson, I.A., Hanke, T., Sliepen, K., Sanders, R.W.(2024) NPJ Vaccines 9: 74-74
- PubMed: 38582771 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41541-024-00862-8
- Primary Citation Related Structures:
8TGO - PubMed Abstract:
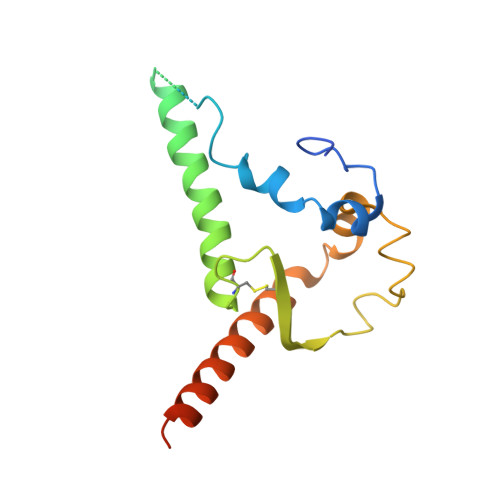
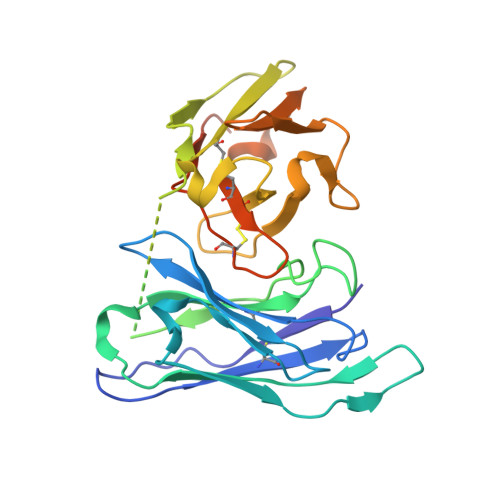
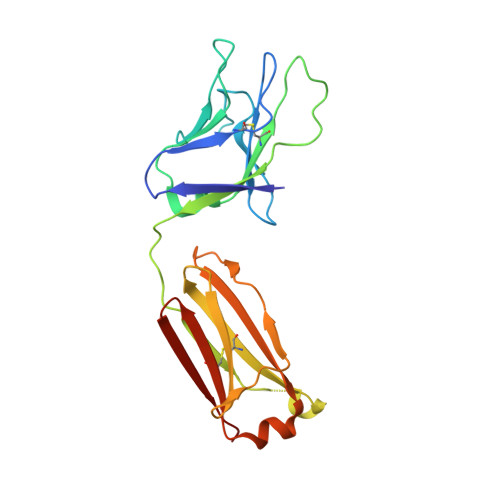
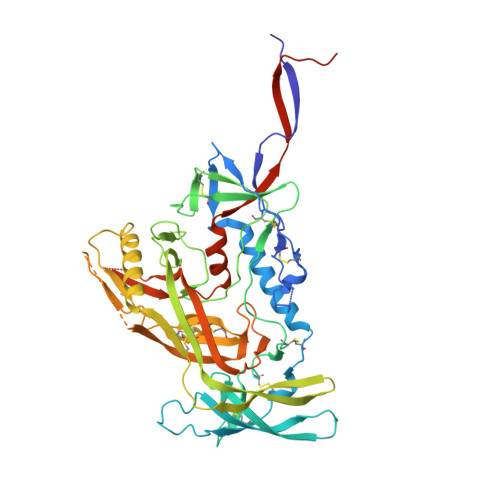
Recombinant native-like HIV-1 envelope glycoprotein (Env) trimers are used in candidate vaccines aimed at inducing broadly neutralizing antibodies. While state-of-the-art SOSIP or single-chain Env designs can be expressed as native-like trimers, undesired monomers, dimers and malformed trimers that elicit non-neutralizing antibodies are also formed, implying that these designs could benefit from further modifications for gene-based vaccination approaches. Here, we describe the triple tandem trimer (TTT) design, in which three Env protomers are genetically linked in a single open reading frame and express as native-like trimers. Viral vectored Env TTT induced similar neutralization titers but with a higher proportion of trimer-specific responses. The TTT design was also applied to generate influenza hemagglutinin (HA) trimers without the need for trimerization domains. Additionally, we used TTT to generate well-folded chimeric Env and HA trimers that harbor protomers from three different strains. In summary, the TTT design is a useful platform for the design of HIV-1 Env and influenza HA immunogens for a multitude of vaccination strategies.
- Department of Medical Microbiology and Infection Prevention, Amsterdam UMC, University of Amsterdam, Amsterdam, Netherlands.
Organizational Affiliation: