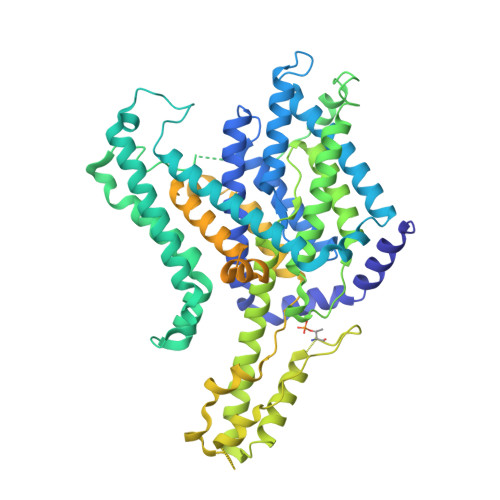
The Bor1 elevator transport cycle is subject to autoinhibition and activation.
Jiang, Y., Jiang, J.(2024) Nat Commun 15: 9090-9090
- PubMed: 39433547 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-53411-1
- Primary Citation Related Structures:
8TEG, 8TEH, 8TEI, 8TEJ, 8TEL, 8TEM, 8TEN - PubMed Abstract:
Boron, essential for plant growth, necessitates precise regulation due to its potential toxicity. This regulation is achieved by borate transporters (BORs), which are homologous to the SLC4 family. The Arabidopsis thaliana Bor1 (AtBor1) transporter from clade I undergoes slow regulation through degradation and translational suppression, but its potential for fast regulation via direct activity modulation was unclear. Here, we combine cryo-electron microscopy, mutagenesis, and functional characterization to study AtBor1, revealing high-resolution structures of the dimer in one inactive and three active states. Our findings show that AtBor1 is regulated by two distinct mechanisms: an autoinhibitory domain at the carboxyl terminus obstructs the substrate pathway via conserved salt bridges, and phosphorylation of Thr410 allows interaction with a positively charged pocket at the cytosolic face, essential for borate transport. These results elucidate the molecular basis of AtBor1's activity regulation and highlight its role in fast boron level regulation in plants.
- Laboratory of Membrane Proteins and Structural Biology, Biochemistry and Biophysics Center, National Heart, Lung, and Blood Institute, National Institutes of Health, Bethesda, MD, USA. yan.jiang@sydney.edu.au.
Organizational Affiliation: