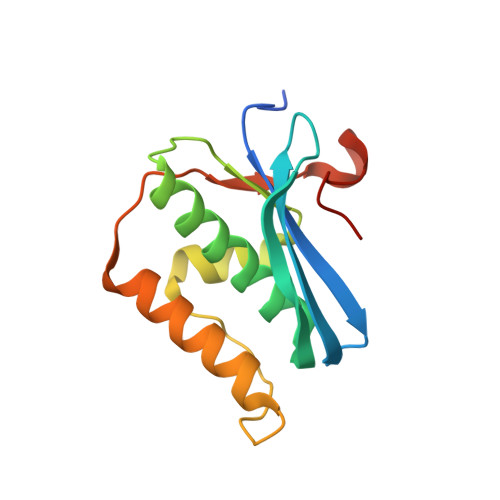
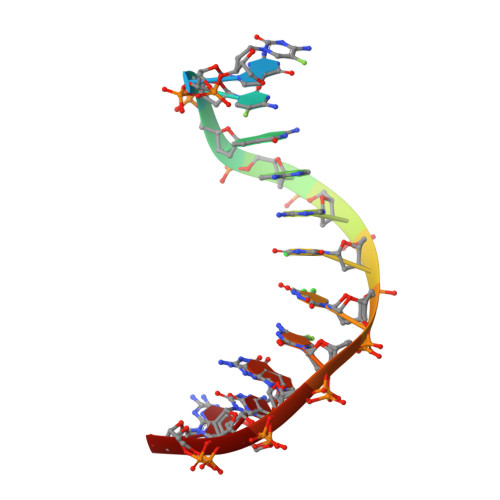
Conformational Morphing by a DNA Analogue Featuring 7-Deazapurines and 5-Halogenpyrimidines and the Origins of Adenine-Tract Geometry.
Pallan, P.S., Lybrand, T.P., Rozners, E., Abramov, M., Schepers, G., Eremeeva, E., Herdewijn, P., Egli, M.(2023) Biochemistry 62: 2854-2867
- PubMed: 37694722 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.3c00327
- Primary Citation Related Structures:
8SV3, 8SV4 - PubMed Abstract:
Several efforts are currently directed at the creation and cellular implementation of alternative genetic systems composed of pairing components that are orthogonal to the natural dA/dT and dG/dC base pairs. In an alternative approach, Watson-Crick-type pairing is conserved, but one or all of the four letters of the A, C, G, and T alphabet are substituted by modified components. Thus, all four nucleobases were altered to create halogenated deazanucleic acid (DZA): dA was replaced by 7-deaza-2'-deoxyadenosine (dzA), dG by 7-deaza-2'-deoxyguanosine (dzG), dC by 5-fluoro-2'-deoxycytidine (FdC), and dT by 5-chloro-2'-deoxyuridine (CldU). This base-pairing system was previously shown to retain function in Escherichia coli . Here, we analyze the stability, hydration, structure, and dynamics of a DZA Dickerson-Drew Dodecamer (DDD) of sequence 5'-FdC-dzG-FdC-dzG-dzA-dzA-CldU-CldU-FdC-dzG-FdC-dzG-3'. Contrary to similar stabilities of DDD and DZA-DDD, osmotic stressing revealed a dramatic loss of hydration for the DZA-DDD relative to that for the DDD. The parent DDD 5'-d(CGCGAATTCGCG)-3' features an A-tract, a run of adenosines uninterrupted by a TpA step, and exhibits a hallmark narrow minor groove. Crystal structures─in the presence of RNase H─and MD simulations show increased conformational plasticity ("morphing") of DZA-DDD relative to that of the DDD. The narrow dzA-tract minor groove in one structure widens to resemble that in canonical B-DNA in a second structure. These changes reflect an indirect consequence of altered DZA major groove electrostatics (less negatively polarized compared to that in DNA) and hydration (reduced compared to that in DNA). Therefore, chemical modifications outside the minor groove that lead to collapse of major groove electrostatics and hydration can modulate A-tract geometry.
- School of Medicine, Department of Biochemistry, and Center for Structural Biology, Vanderbilt University, Nashville, Tennessee 37232, United States.
Organizational Affiliation: