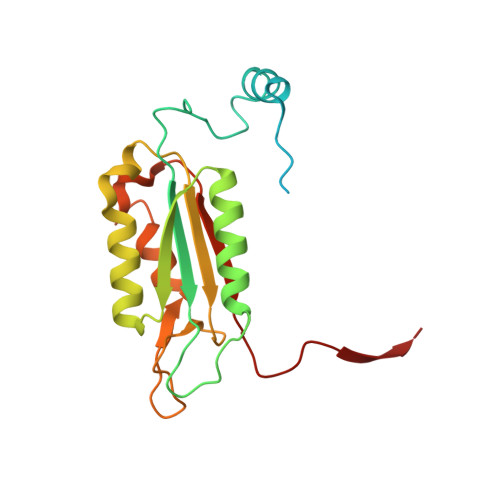
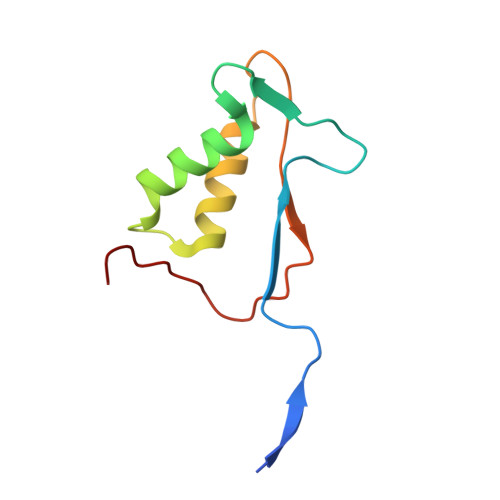
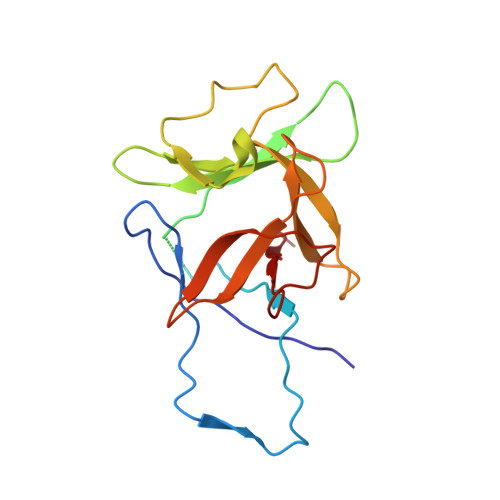
Structural insights into cytokine cleavage by inflammatory caspase-4.
Devant, P., Dong, Y., Mintseris, J., Ma, W., Gygi, S.P., Wu, H., Kagan, J.C.(2023) Nature 624: 451-459
- PubMed: 37993712 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-023-06751-9
- Primary Citation Related Structures:
8SPB - PubMed Abstract:
Inflammatory caspases are key enzymes in mammalian innate immunity that control the processing and release of interleukin-1 (IL-1)-family cytokines 1,2 . Despite the biological importance, the structural basis for inflammatory caspase-mediated cytokine processing has remained unclear. To date, catalytic cleavage of IL-1-family members, including pro-IL-1β and pro-IL-18, has been attributed primarily to caspase-1 activities within canonical inflammasomes 3 . Here we demonstrate that the lipopolysaccharide receptor caspase-4 from humans and other mammalian species (except rodents) can cleave pro-IL-18 with an efficiency similar to pro-IL-1β and pro-IL-18 cleavage by the prototypical IL-1-converting enzyme caspase-1. This ability of caspase-4 to cleave pro-IL-18, combined with its previously defined ability to cleave and activate the lytic pore-forming protein gasdermin D (GSDMD) 4,5 , enables human cells to bypass the need for canonical inflammasomes and caspase-1 for IL-18 release. The structure of the caspase-4-pro-IL-18 complex determined using cryogenic electron microscopy reveals that pro-lL-18 interacts with caspase-4 through two distinct interfaces: a protease exosite and an interface at the caspase-4 active site involving residues in the pro-domain of pro-IL-18, including the tetrapeptide caspase-recognition sequence 6 . The mechanisms revealed for cytokine substrate capture and cleavage differ from those observed for the caspase substrate GSDMD 7,8 . These findings provide a structural framework for the discussion of caspase activities in health and disease.
- Division of Gastroenterology, Boston Children's Hospital and Harvard Medical School, Boston, MA, USA.
Organizational Affiliation: