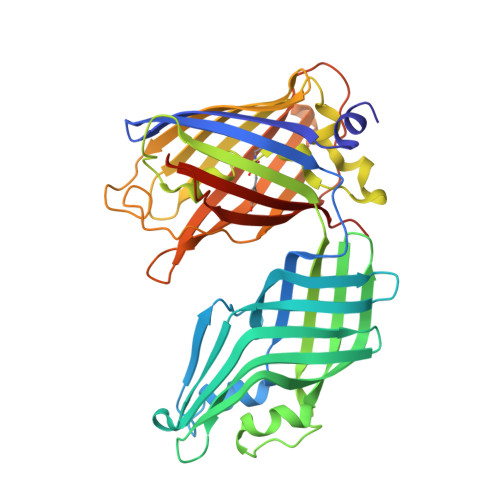
Development and atomic structure of a new fluorescence-based sensor to probe heme transfer in bacterial pathogens.
Mahoney, B.J., Goring, A.K., Wang, Y., Dasika, P., Zhou, A., Grossbard, E., Cascio, D., Loo, J.A., Clubb, R.T.(2023) J Inorg Biochem 249: 112368-112368
- PubMed: 37729854 Search on PubMed
- DOI: https://doi.org/10.1016/j.jinorgbio.2023.112368
- Primary Citation Related Structures:
8SMU - PubMed Abstract:
Heme is the most abundant source of iron in the human body and is actively scavenged by bacterial pathogens during infections. Corynebacterium diphtheriae and other species of actinobacteria scavenge heme using cell wall associated and secreted proteins that contain Conserved Region (CR) domains. Here we report the development of a fluorescent sensor to measure heme transfer from the C-terminal CR domain within the HtaA protein (CR2) to other hemoproteins within the heme-uptake system. The sensor contains the CR2 domain inserted into the β2 to β3 turn of the Enhanced Green Fluorescent Protein (EGFP). A 2.45 Å crystal structure reveals the basis of heme binding to the CR2 domain via iron-tyrosyl coordination and shares conserved structural features with CR domains present in Corynebacterium glutamicum. The structure and small angle X-ray scattering experiments are consistent with the sensor adopting a V-shaped structure that exhibits only small fluctuations in inter-domain positioning. We demonstrate heme transfer from the sensor to the CR domains located within the HtaA or HtaB proteins in the heme-uptake system as measured by a ∼ 60% increase in sensor fluorescence and native mass spectrometry.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, 611 Charles E. Young Drive East, Los Angeles, CA 90095, USA; UCLA-DOE Institute of Genomics and Proteomics, University of California, Los Angeles, 611 Charles E. Young Drive East, Los Angeles, CA 90095, USA.
Organizational Affiliation: