In silico lambda-dynamics predicts protein binding specificities to modified RNAs.
Angelo, M., Zhang, W., Vilseck, J.Z., Aoki, S.T.(2025) Nucleic Acids Res 53
- PubMed: 40066880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaf166
- Primary Citation Related Structures:
8SIP, 8TCA, 8VEV - PubMed Abstract:
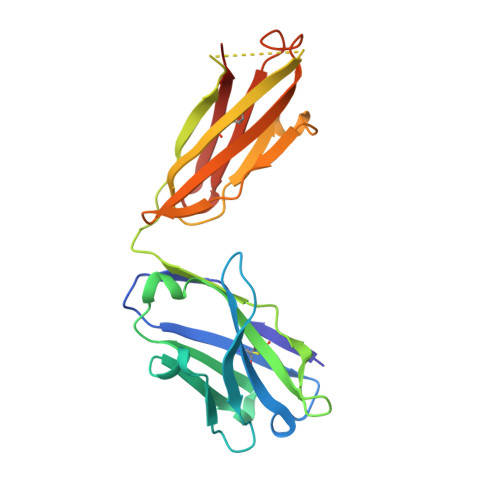
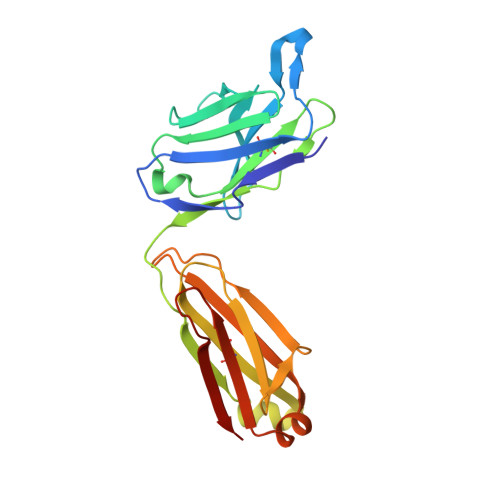
RNA modifications shape gene expression through a variety of chemical changes to canonical RNA bases. Although numbering in the hundreds, only a few RNA modifications are well characterized, in part due to the absence of methods to identify modification sites. Antibodies remain a common tool to identify modified RNA and infer modification sites through straightforward applications. However, specificity issues can result in off-target binding and confound conclusions. This work utilizes in silico λ-dynamics to efficiently estimate binding free energy differences of modification-targeting antibodies between a variety of naturally occurring RNA modifications. Crystal structures of inosine and N6-methyladenosine (m6A) targeting antibodies bound to their modified ribonucleosides were determined and served as structural starting points. λ-Dynamics was utilized to predict RNA modifications that permit or inhibit binding to these antibodies. In vitro RNA-antibody binding assays supported the accuracy of these in silico results. High agreement between experimental and computed binding propensities demonstrated that λ-dynamics can serve as a predictive screen for antibody specificity against libraries of RNA modifications. More importantly, this strategy is an innovative way to elucidate how hundreds of known RNA modifications interact with biological molecules without the limitations imposed by in vitro or in vivo methodologies.
- Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, 635 Barnhill Drive, Indianapolis, IN 46202, United States.
Organizational Affiliation: