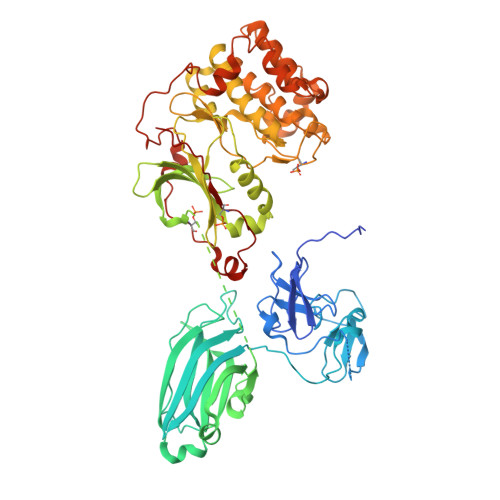
Molecular Basis of Allosteric Regulation and Pharmaceutical Targeting of Protein Kinase C b
Cong, A.T.Q., Witter, T.L., Bruinsma, E.S., Bhattacharya, S.S., Jayaraman, S., Dugan, M.B., Paluncic, J., Kuffel, M.J., Farmakes, J., Alvey, J., Wu, X., Fields, A.P., Pandey, A., Hawse, J.R., Goetz, M.P., Schellenberg, M.J.To be published.