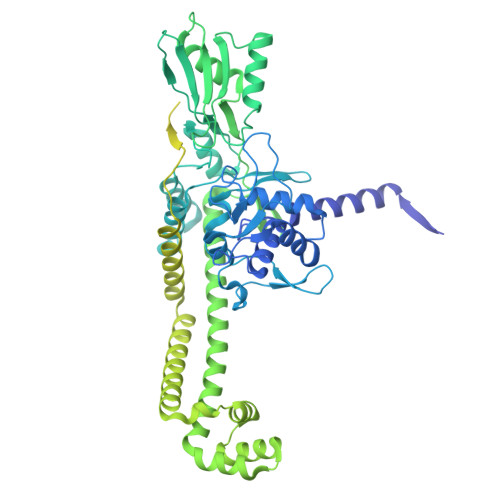
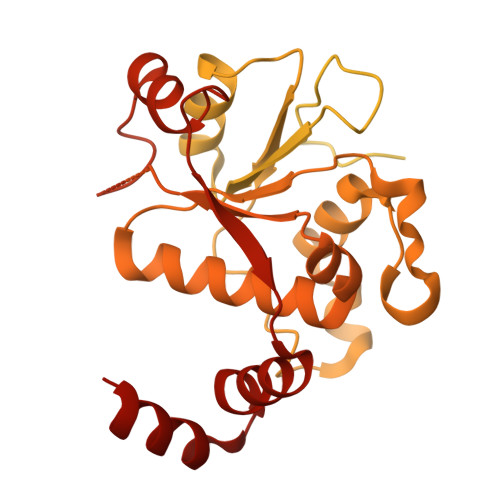
Molecular mechanism of a triazole-containing inhibitor of Mycobacterium tuberculosis DNA gyrase.
Gedeon, A., Yab, E., Dinut, A., Sadowski, E., Capton, E., Dreneau, A., Petit, J., Gioia, B., Piveteau, C., Djaout, K., Lecat, E., Wehenkel, A.M., Gubellini, F., Mechaly, A., Alzari, P.M., Deprez, B., Baulard, A., Aubry, A., Willand, N., Petrella, S.(2024) iScience 27: 110967-110967
- PubMed: 39429773 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2024.110967
- Primary Citation Related Structures:
8S7K, 8S7O - PubMed Abstract:
Antimicrobial resistance remains a persistent and pressing public health concern. Here, we describe the synthesis of original triazole-containing inhibitors targeting the DNA gyrase, a well-validated drug target for developing new antibiotics. Our compounds demonstrate potent antibacterial activity against various pathogenic bacteria, with notable potency against Mycobacterium tuberculosis ( Mtb ). Moreover, one hit, compound 10a , named BDM71403, was shown to be more potent in Mtb than the NBTI of reference, gepotidacin. Mechanistic enzymology assays reveal a competitive interaction of BDM71403 with fluoroquinolones within the Mtb gyrase cleavage core. High-resolution cryo-electron microscopy structural analysis provides detailed insights into the ternary complex formed by the Mtb gyrase, double-stranded DNA, and either BDM71403 or gepotidacin, providing a rational framework to understand the superior in vitro efficacy on Mtb . This study highlights the potential of triazole-based scaffolds as promising gyrase inhibitors, offering new avenues for drug development in the fight against antimicrobial resistance.
- Institut Pasteur, Université Paris Cité, CNRS UMR 3528, Unité de Microbiologie Structurale, 75015 Paris, France.
Organizational Affiliation: