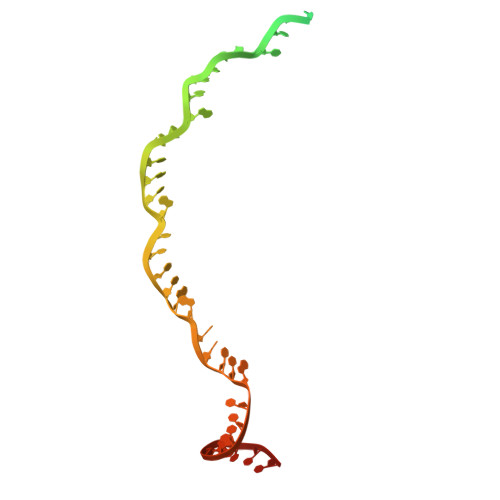
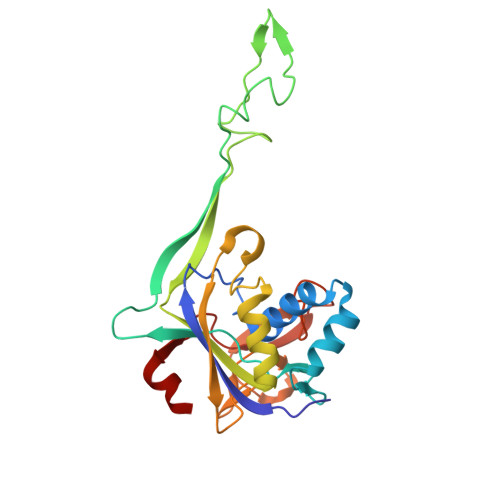
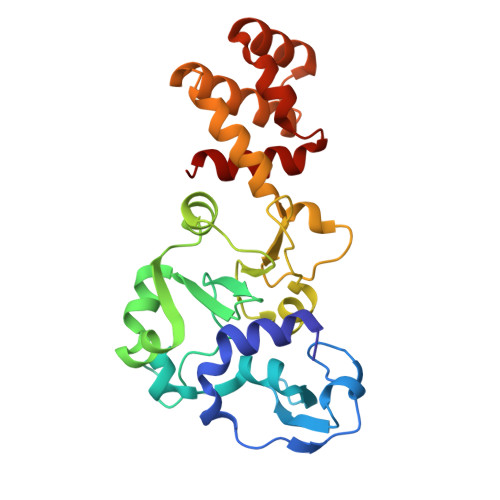
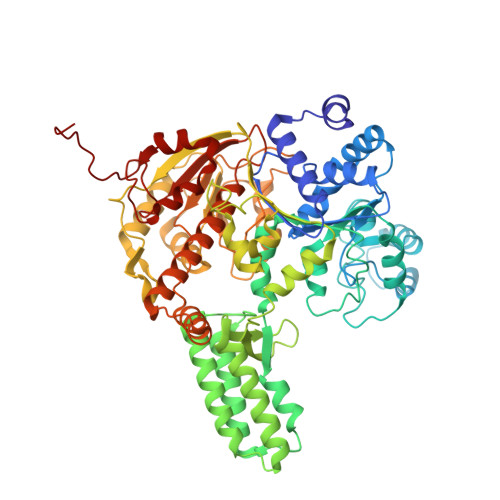
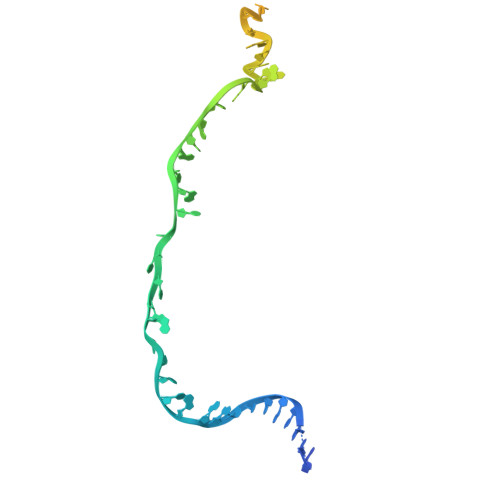
Structural variation of types IV-A1- and IV-A3-mediated CRISPR interference.
Cepaite, R., Klein, N., Miksys, A., Camara-Wilpert, S., Ragozius, V., Benz, F., Skorupskaite, A., Becker, H., Zvejyte, G., Steube, N., Hochberg, G.K.A., Randau, L., Pinilla-Redondo, R., Malinauskaite, L., Pausch, P.(2024) Nat Commun 15: 9306-9306
- PubMed: 39468082 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-53778-1
- Primary Citation Related Structures:
8RC2, 8RC3, 8RFJ, 8S35, 8S36, 8S37 - PubMed Abstract:
CRISPR-Cas mediated DNA-interference typically relies on sequence-specific binding and nucleolytic degradation of foreign genetic material. Type IV-A CRISPR-Cas systems diverge from this general mechanism, using a nuclease-independent interference pathway to suppress gene expression for gene regulation and plasmid competition. To understand how the type IV-A system associated effector complex achieves this interference, we determine cryo-EM structures of two evolutionarily distinct type IV-A complexes (types IV-A1 and IV-A3) bound to cognate DNA-targets in the presence and absence of the type IV-A signature DinG effector helicase. The structures reveal how the effector complexes recognize the protospacer adjacent motif and target-strand DNA to form an R-loop structure. Additionally, we reveal differences between types IV-A1 and IV-A3 in DNA interactions and structural motifs that allow for in trans recruitment of DinG. Our study provides a detailed view of type IV-A mediated DNA-interference and presents a structural foundation for engineering type IV-A-based genome editing tools.
- LSC-EMBL Partnership Institute for Genome Editing Technologies, Life Sciences Center, Vilnius University, Vilnius, Lithuania.
Organizational Affiliation: