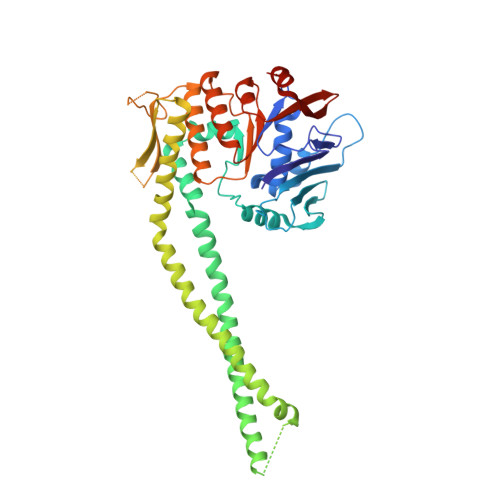
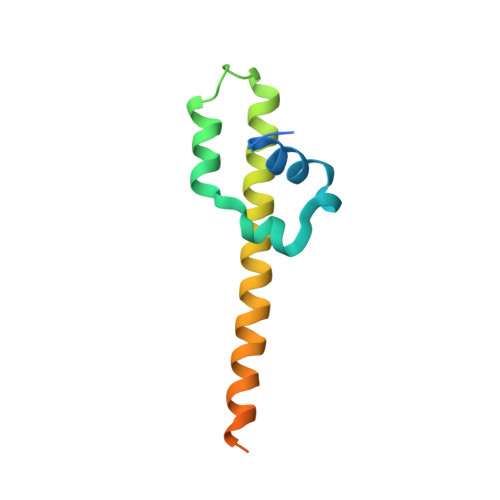
The cohesin ATPase cycle is mediated by specific conformational dynamics and interface plasticity of SMC1A and SMC3 ATPase domains.
Vitoria Gomes, M., Landwerlin, P., Diebold-Durand, M.L., Shaik, T.B., Durand, A., Troesch, E., Weber, C., Brillet, K., Lemee, M.V., Decroos, C., Dulac, L., Antony, P., Watrin, E., Ennifar, E., Golzio, C., Romier, C.(2024) Cell Rep 43: 114656-114656
- PubMed: 39240714 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2024.114656
- Primary Citation Related Structures:
8P0A, 8PQ5, 8RO6, 8RO7, 8RO8, 8RO9, 8ROA, 8ROB, 8ROC, 8ROD, 8ROE, 8ROF, 8ROG, 8ROH, 8ROI, 8ROJ, 8ROK, 8ROL - PubMed Abstract:
Cohesin is key to eukaryotic genome organization and acts throughout the cell cycle in an ATP-dependent manner. The mechanisms underlying cohesin ATPase activity are poorly understood. Here, we characterize distinct steps of the human cohesin ATPase cycle and show that the SMC1A and SMC3 ATPase domains undergo specific but concerted structural rearrangements along this cycle. Specifically, whereas the proximal coiled coil of the SMC1A ATPase domain remains conformationally stable, that of the SMC3 displays an intrinsic flexibility. The ATP-dependent formation of the heterodimeric SMC1A/SMC3 ATPase module (engaged state) favors this flexibility, which is counteracted by NIPBL and DNA binding (clamped state). Opening of the SMC3/RAD21 interface (open-engaged state) stiffens the SMC3 proximal coiled coil, thus constricting together with that of SMC1A the ATPase module DNA-binding chamber. The plasticity of the ATP-dependent interface between the SMC1A and SMC3 ATPase domains enables these structural rearrangements while keeping the ATP gate shut. VIDEO ABSTRACT.
- Université de Strasbourg, IGBMC UMR 7104 - UMR-S 1258, 67400 Illkirch, France; CNRS, UMR 7104, 67400 Illkirch, France; INSERM, UMR-S 1258, 67400 Illkirch, France; Institut de Génétique et de Biologie Moléculaire et Cellulaire, Department of Integrated Structural Biology, 67400 Illkirch, France.
Organizational Affiliation: