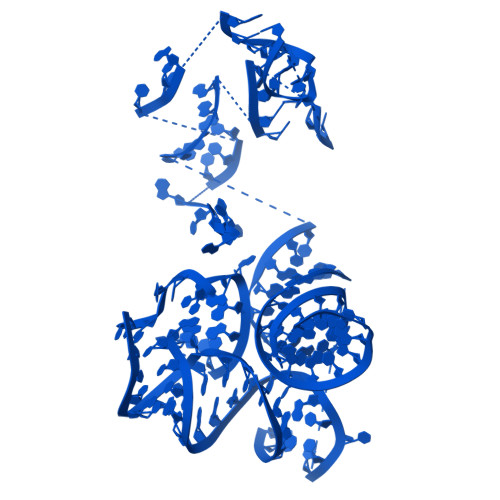
Structural basis for translational control by the human 48S initiation complex.
Petrychenko, V., Yi, S.H., Liedtke, D., Peng, B.Z., Rodnina, M.V., Fischer, N.(2025) Nat Struct Mol Biol 32: 62-72
- PubMed: 39289545 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01378-4
- Primary Citation Related Structures:
8PJ1, 8PJ2, 8PJ3, 8PJ4, 8PJ5, 8PJ6, 8RG0 - PubMed Abstract:
The selection of an open reading frame (ORF) for translation of eukaryotic mRNA relies on remodeling of the scanning 48S initiation complex into an elongation-ready 80S ribosome. Using cryo-electron microscopy, we visualize the key commitment steps orchestrating 48S remodeling in humans. The mRNA Kozak sequence facilitates mRNA scanning in the 48S open state and stabilizes the 48S closed state by organizing the contacts of eukaryotic initiation factors (eIFs) and ribosomal proteins and by reconfiguring mRNA structure. GTPase-triggered large-scale fluctuations of 48S-bound eIF2 facilitate eIF5B recruitment, transfer of initiator tRNA from eIF2 to eIF5B and the release of eIF5 and eIF2. The 48S-bound multisubunit eIF3 complex controls ribosomal subunit joining by coupling eIF exchange to gradual displacement of the eIF3c N-terminal domain from the intersubunit interface. These findings reveal the structural mechanism of ORF selection in human cells and explain how eIF3 could function in the context of the 80S ribosome.
- Project Group Molecular Machines in Motion, Department of Physical Biochemistry, Max Planck Institute for Multidisciplinary Sciences, Göttingen, Germany.
Organizational Affiliation: