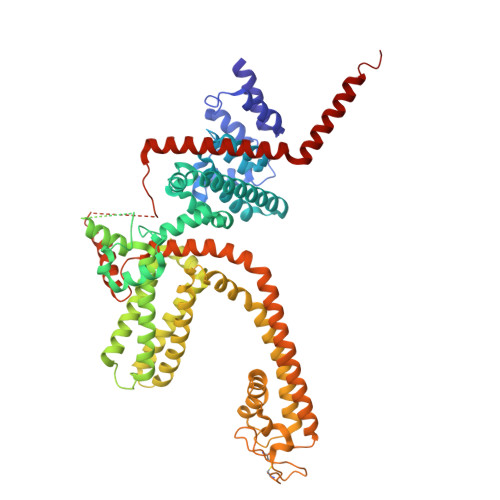
Identification of a binding site for small molecule inhibitors targeting human TRPM4.
Ekundayo, B., Arullampalam, P., Gerber, C.E., Hammerli, A.F., Guichard, S., Boukenna, M., Ross-Kaschitza, D., Lochner, M., Rougier, J.S., Stahlberg, H., Abriel, H., Ni, D.(2025) Nat Commun 16: 833-833
- PubMed: 39828793 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-56131-2
- Primary Citation Related Structures:
8RCR, 8RCU, 8RD9 - PubMed Abstract:
Transient receptor potential (TRP) melastatin 4 (TRPM4) protein is a calcium-activated monovalent cation channel associated with various genetic and cardiovascular disorders. The anthranilic acid derivative NBA is a potent and specific TRPM4 inhibitor, but its binding site in TRPM4 has been unknown, although this information is crucial for drug development targeting TRPM4. We determine three cryo-EM structures of full-length human TRPM4 embedded in native lipid nanodiscs without inhibitor, bound to NBA, and an anthranilic acid derivative, IBA. We found that the small molecules NBA and IBA were bound in a pocket formed between the S3, S4, and TRP helices and the S4-S5 linker of TRPM4. Our structural data and results from patch clamp experiments enable validation of a binding site for small molecule inhibitors, paving the way for further drug development targeting TRPM4.
- Laboratory of Biological Electron Microscopy, IPHYS, SB, EPFL, and Dept. Fundamental Microbiology, Faculty of Biology and Medicine, UNIL, Cubotron, Rt. de la Sorge, Lausanne, Switzerland.
Organizational Affiliation: