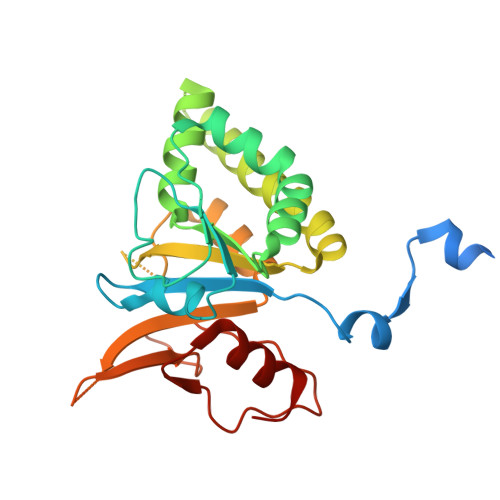
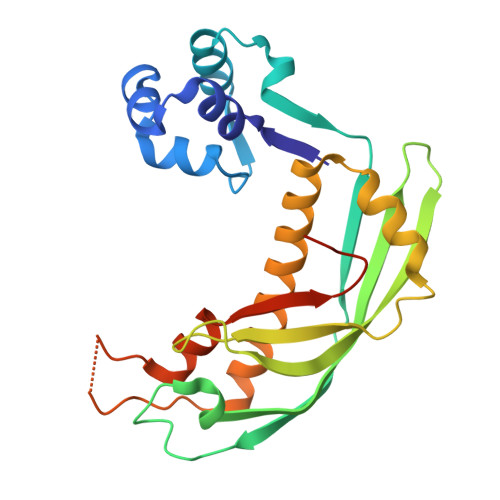
Phage parasites targeting phage homologous recombinases provide antiviral immunity.
Debiasi-Anders, G., Qiao, C., Salim, A., Li, N., Mir-Sanchis, I.(2025) Nat Commun 16: 1889-1889
- PubMed: 39987160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-57156-3
- Primary Citation Related Structures:
8PQ8, 8Q86, 8QE9, 8RC5 - PubMed Abstract:
Bacteria often carry multiple genes encoding anti-phage defense systems, clustered in defense islands and phage satellites. Various unrelated anti-phage defense systems target phage-encoded homologous recombinases (HRs) through unclear mechanisms. Here, we show that the phage satellite SaPI2, which does not encode orthodox anti-phage defense systems, provides antiviral immunity mediated by Stl2, the SaPI2-encoded transcriptional repressor. Stl2 targets and inhibits phage-encoded HRs, including Sak and Sak4, two HRs from the Rad52-like and Rad51-like superfamilies. Remarkably, apo Stl2 forms a collar of dimers oligomerizing as closed rings and as filaments, mimicking the quaternary structure of its targets. Stl2 decorates both Sak rings and Sak4 filaments. The oligomerization of Stl2 as a collar of dimers is necessary for its inhibitory activity both in vitro and in vivo. Our results shed light on the mechanisms underlying antiviral immunity against phages carrying divergent HRs.
- Department of Medical Biochemistry and Biophysics, Umeå University, Umeå, Sweden.
Organizational Affiliation: