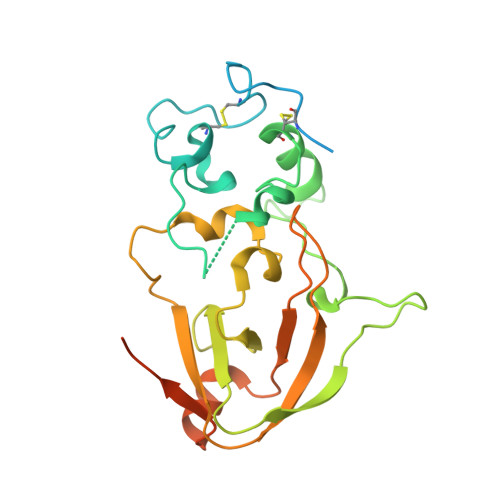
Evolution of fungal tuberculosis necrotizing toxin (TNT) domain-containing enzymes reveals divergent adaptations to enhance NAD cleavage.
Ferrario, E., Kallio, J.P., Emdadi, M., Stromland, O., Rack, J.G.M., Ziegler, M.(2024) Protein Sci 33: e5071-e5071
- PubMed: 38895984 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5071
- Primary Citation Related Structures:
8R15, 8R17 - PubMed Abstract:
Tuberculosis necrotizing toxin (TNT) is a protein domain discovered on the outer membrane of Mycobacterium tuberculosis (Mtb), and the fungal pathogen Aspergillus fumigatus. TNT domains have pure NAD(P) hydrolytic activity, setting them apart from other NAD-cleaving domains such as ADP-ribosyl cyclase and Toll/interleukin-1 receptor homology (TIR) domains which form a wider set of products. Importantly, the Mtb TNT domain has been shown to be involved in immune evasion via depletion of the intracellular NAD pool of macrophages. Therefore, an intriguing hypothesis is that TNT domains act as "NAD killers" in host cells facilitating pathogenesis. Here, we explore the phylogenetic distribution of TNT domains and detect their presence solely in bacteria and fungi. Within fungi, we discerned six TNT clades. In addition, X-ray crystallography and AlphaFold2 modeling unveiled clade-specific strategies to promote homodimer stabilization of the fungal enzymes, namely, Ca 2+ binding, disulfide bonds, or hydrogen bonds. We show that dimer stabilization is a requirement for NADase activity and that the group-specific strategies affect the active site conformation, thereby modulating enzyme activity. Together, these findings reveal the evolutionary lineage of fungal TNT enzymes, corroborating the hypothesis of them being pure extracellular NAD (eNAD) cleavers, with possible involvement in microbial warfare and host immune evasion.
- Department of Biomedicine, University of Bergen, Bergen, Norway.
Organizational Affiliation: