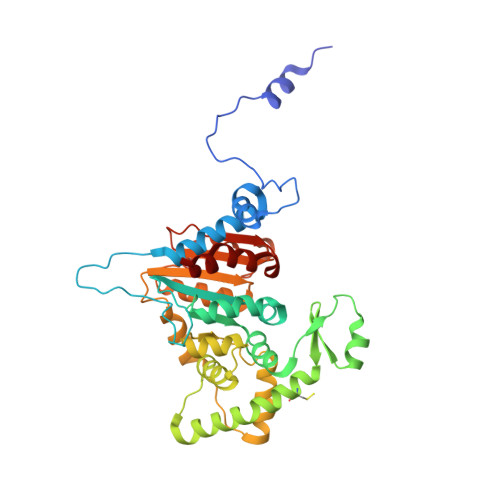
Structural characterization of the (deoxy)hypusination in Trichomonas vaginalis questions the bifunctionality of deoxyhypusine synthase.
Wator, E., Wilk, P., Kochanowski, P., Grudnik, P.(2024) FEBS J 291: 3856-3869
- PubMed: 38923395 Search on PubMed
- DOI: https://doi.org/10.1111/febs.17207
- Primary Citation Related Structures:
8QZV, 8QZW, 8QZX - PubMed Abstract:
Trichomonas vaginalis, the causative agent of trichomoniasis, is a prevalent anaerobic protozoan parasite responsible for the most common nonviral sexually transmitted infection globally. While metronidazole and its derivatives are approved drugs for this infection, rising resistance necessitates the exploration of new antiparasitic therapies. Protein posttranslational modifications (PTMs) play crucial roles in cellular processes, and among them, hypusination, involving eukaryotic translation factor 5A (eIF5A), has profound implications. Despite extensive studies in various organisms, the role of hypusination in T. vaginalis and its potential impact on parasite biology and pathogenicity remain poorly understood. This study aims to unravel the structural basis of the hypusination pathway in T. vaginalis using X-ray crystallography and cryo-electron microscopy. The results reveal high structural homology between T. vaginalis and human orthologs, providing insights into the molecular architecture of eIF5A and deoxyhypusine synthase (DHS) and their interaction. Contrary to previous suggestions of bifunctionality, our analyses indicate that the putative hydroxylation site in tvDHS is nonfunctional, and biochemical assays demonstrate exclusive deoxyhypusination capability. These findings challenge the notion of tvDHS functioning as both deoxyhypusine synthase and hydroxylase. The study enhances understanding of the hypusination pathway in T. vaginalis, shedding light on its functional relevance and potential as a drug target, and contributing to the development of novel therapeutic strategies against trichomoniasis.
- Małopolska Centre of Biotechnology, Jagiellonian University, Kraków, Poland.
Organizational Affiliation: