Exploring sequence space and engineering of Xpo1-dependent NESes
Rymarenko, O., Huyton, T., Gorlich, D.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| GTP-binding nuclear protein GSP1/CNR1 | A, C [auth B], E, G | 182 | Saccharomyces cerevisiae S288C | Mutation(s): 1 Gene Names: GSP1 | 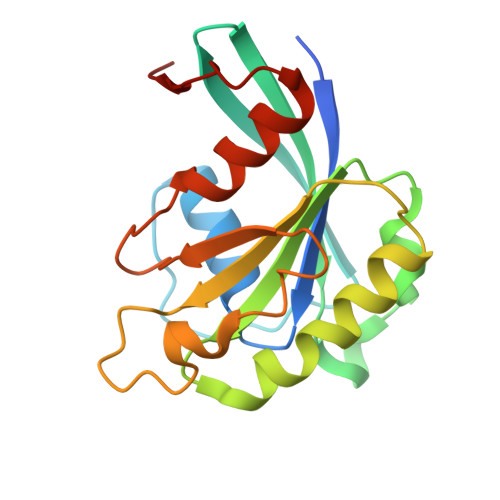 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P32835 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Exportin-1 | B [auth C], D, F, H | 1,048 | Saccharomyces cerevisiae S288C | Mutation(s): 0 Gene Names: CRM1 | 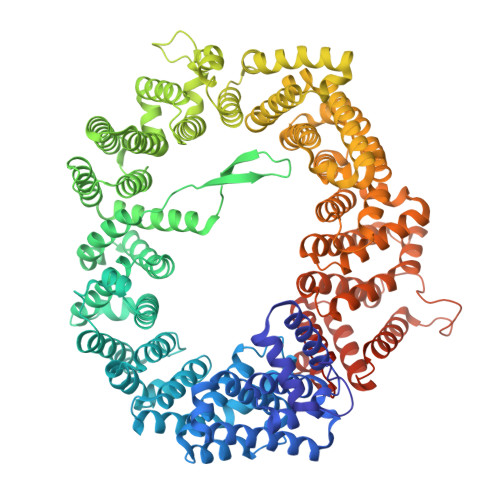 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P30822 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| hiNES2 | 16 | synthetic construct | Mutation(s): 0 |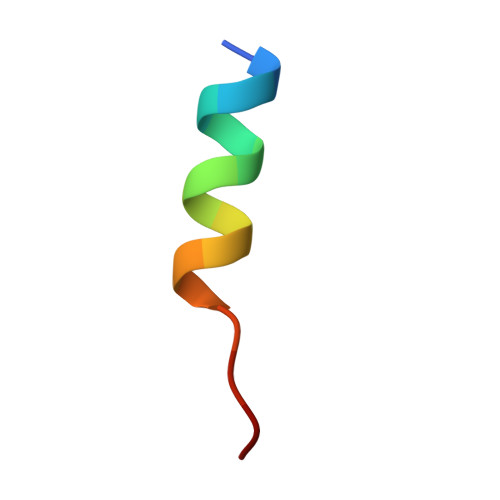  | |
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GTP Download:Ideal Coordinates CCD File | M [auth A], P [auth B], S [auth E], V [auth G] | GUANOSINE-5'-TRIPHOSPHATE C10 H16 N5 O14 P3 XKMLYUALXHKNFT-UUOKFMHZSA-N |  | ||
| ACT Download:Ideal Coordinates CCD File | O [auth A], R [auth B], U [auth E], X [auth G] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| MG Download:Ideal Coordinates CCD File | N [auth A], Q [auth B], T [auth E], W [auth G] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 97.355 | α = 82.08 |
| b = 105.572 | β = 86.668 |
| c = 170.4 | γ = 76.668 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| Coot | model building |
| DIALS | data reduction |
| DIALS | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Max Planck Society | Germany | -- |