Bioinformatic, Enzymatic, and Structural Characterization of Trichuris suis Hexosaminidase HEX-2.
Dutkiewicz, Z., Varrot, A., Breese, K.J., Stubbs, K.A., Nuschy, L., Adduci, I., Paschinger, K., Wilson, I.B.H.(2024) Biochemistry 63: 1941-1954
- PubMed: 39058279 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.4c00187
- Primary Citation Related Structures:
8QK1 - PubMed Abstract:
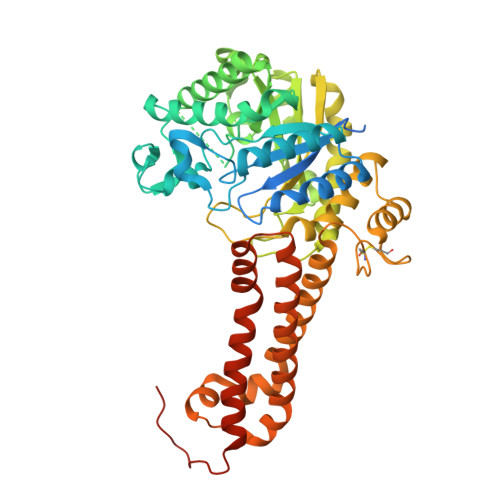
Hexosaminidases are key enzymes in glycoconjugate metabolism and occur in all kingdoms of life. Here, we have investigated the phylogeny of the GH20 glycosyl hydrolase family in nematodes and identified a β-hexosaminidase subclade present only in the Dorylaimia. We have expressed one of these, HEX-2 from Trichuris suis , a porcine parasite, and shown that it prefers an aryl β- N -acetylgalactosaminide in vitro . HEX-2 has an almost neutral pH optimum and is best inhibited by GalNAc-isofagomine. Toward N-glycan substrates, it displays a preference for the removal of GalNAc residues from LacdiNAc motifs as well as the GlcNAc attached to the α1,3-linked core mannose. Therefore, it has a broader specificity than insect fused lobe (FDL) hexosaminidases but one narrower than distant homologues from plants. Its X-ray crystal structure, the first of any subfamily 1 GH20 hexosaminidase to be determined, is closest to Streptococcus pneumoniae GH20C and the active site is predicted to be compatible with accommodating both GalNAc and GlcNAc. The new structure extends our knowledge about this large enzyme family, particularly as T. suis HEX-2 also possesses the key glutamate residue found in human hexosaminidases of either GH20 subfamily, including HEXD whose biological function remains elusive.
- Institut für Biochemie, Department für Chemie, Universität für Bodenkultur, Muthgasse 18, Wien 1190, Austria.
Organizational Affiliation: