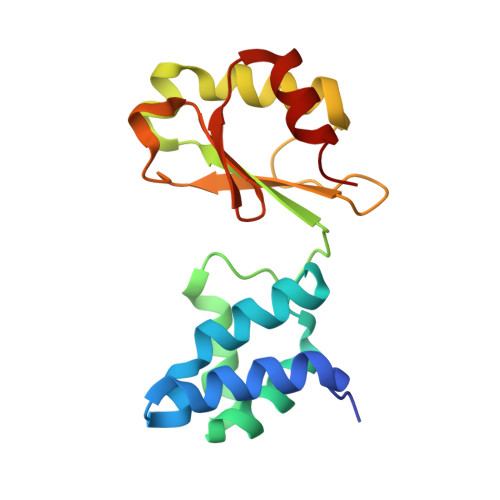
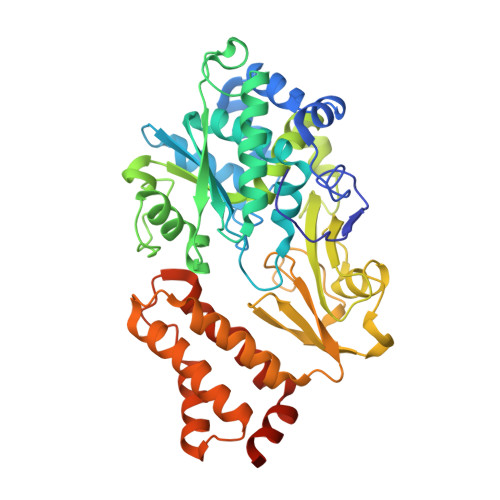
Structures of 3-acetylpyridine adenine dinucleotide and ADP-ribose bound to the electron input module of respiratory complex I.
Wohlwend, D., Merono, L., Bucka, S., Ritter, K., Jessen, H.J., Friedrich, T.(2024) Structure 32: 715
- PubMed: 38503292 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2024.02.013
- Primary Citation Related Structures:
8QG1, 8QGW, 8QH4, 8QH7, 8QHK - PubMed Abstract:
Energy-converting NADH:ubiquinone oxidoreductase, respiratory complex I, is a major enzyme of energy metabolism that couples NADH oxidation and ubiquinone reduction with proton translocation. The NADH oxidation site features different enzymatic activities with various nucleotides. While the kinetics of these reactions are well described, only binding of NAD + and NADH have been structurally characterized. Here, we report the structures of the electron input module of Aquifex aeolicus complex I with bound ADP-ribose and 3-acetylpyridine adenine dinucleotides at resolutions better than 2.0 Å. ADP-ribose acts as inhibitor by blocking the "ADP-handle" motif essential for nucleotide binding. The pyridine group of APADH is minimally offset from flavin, which could contribute to its poorer suitability as substrate. A comparison with other nucleotide co-structures surprisingly shows that the adenine ribose and the pyrophosphate moiety contribute most to nucleotide binding, thus all adenine dinucleotides share core binding modes to the unique Rossmann-fold in complex I.
- Institute of Biochemistry, Albert-Ludwigs-Universität Freiburg, Freiburg, Germany.
Organizational Affiliation: