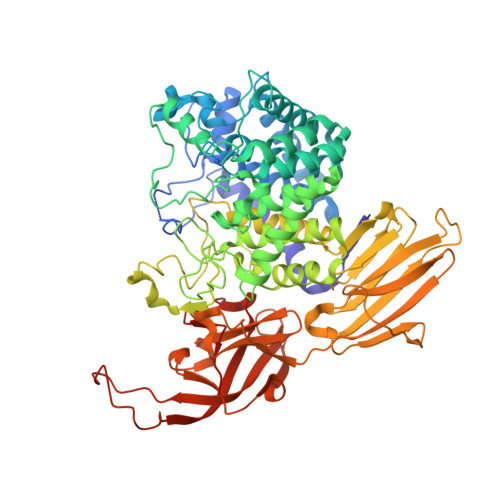
beta-l- Arabino furano-cyclitol Aziridines Are Covalent Broad-Spectrum Inhibitors and Activity-Based Probes for Retaining beta-l-Arabinofuranosidases.
Borlandelli, V., Offen, W., Moroz, O., Nin-Hill, A., McGregor, N., Binkhorst, L., Ishiwata, A., Armstrong, Z., Artola, M., Rovira, C., Davies, G.J., Overkleeft, H.S.(2023) ACS Chem Biol 18: 2564-2573
- PubMed: 38051515 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.3c00558
- Primary Citation Related Structures:
8QF2, 8QF8 - PubMed Abstract:
GH127 and GH146 microorganismal retaining β-l-arabinofuranosidases, expressed by human gut microbiomes, feature an atypical catalytic domain and an unusual mechanism of action. We recently reported that both Bacteroides thetaiotaomicron Bt GH146 and Bifidobacterium longum HypBA1 are inhibited by β-l- arabino furanosyl cyclophellitol epoxide, supporting the action of a zinc-coordinated cysteine as a catalytic nucleophile, where in most retaining GH families, an aspartate or glutamate is employed. This work presents a panel of β-l- arabino furanosyl cyclophellitol epoxides and aziridines as mechanism-based Bt GH146/HypBA1 inhibitors and activity-based probes. The β-l- arabino furanosyl cyclophellitol aziridines both inhibit and label β-l-arabinofuranosidase efficiently (however with different activities), whereas the epoxide-derived probes favor Bt GH146 over HypBA1. These findings are accompanied by X-ray structural analysis of the unmodified β-l- arabino furanosyl cyclophellitol aziridine in complex with both isozymes, which were shown to react by nucleophilic opening of the aziridine, at the pseudoanomeric carbon, by the active site cysteine nucleophile to form a stable thioether bond. Altogether, our activity-based probes may serve as chemical tools for the detection and identification of low-abundance β-l-arabinofuranosidases in complex biological samples.
- Bio-organic Synthesis, Leiden Institute of Chemistry (LIC), Leiden University, Gorlaeus Laboratories, Einsteinweg 55, 2333 CC Leiden, The Netherlands.
Organizational Affiliation: