Crystal structure of apo-GltTk obtained with in meso crystallization (P6322 space group)
Marin, E., Guskov, A., Borshchevskiy, V., Kovalev, K.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Proton/glutamate symporter, SDF family | 430 | Thermococcus kodakarensis KOD1 | Mutation(s): 0 Gene Names: TK0986 Membrane Entity: Yes | 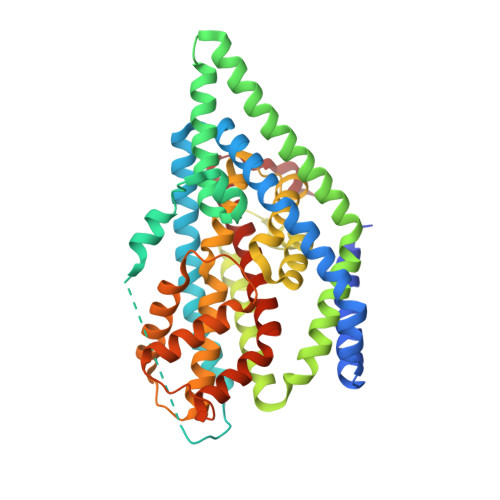 | |
UniProt | |||||
Find proteins for Q5JID0 (Thermococcus kodakarensis (strain ATCC BAA-918 / JCM 12380 / KOD1)) Explore Q5JID0 Go to UniProtKB: Q5JID0 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q5JID0 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| OLC Query on OLC | E [auth A] | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate C21 H40 O4 RZRNAYUHWVFMIP-GDCKJWNLSA-N |  | ||
| OLA Query on OLA | B [auth A], C [auth A], D [auth A], F [auth A] | OLEIC ACID C18 H34 O2 ZQPPMHVWECSIRJ-KTKRTIGZSA-N |  | ||
| SO4 Query on SO4 | G [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 94.24 | α = 90 |
| b = 94.24 | β = 90 |
| c = 244.56 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Netherlands Organisation for Scientific Research (NWO) | Netherlands | OCENW.KLEIN.141 |