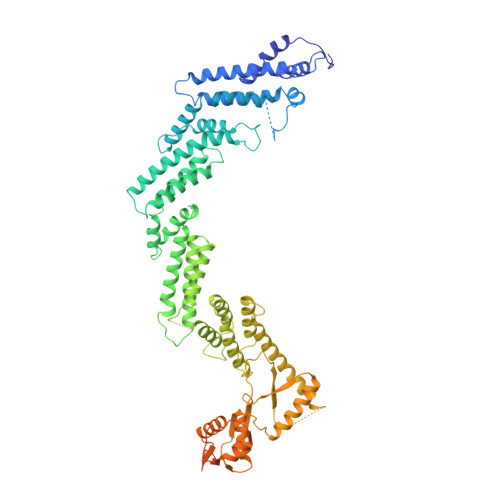
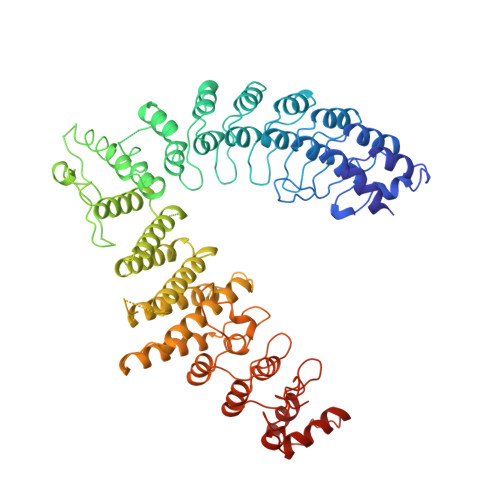
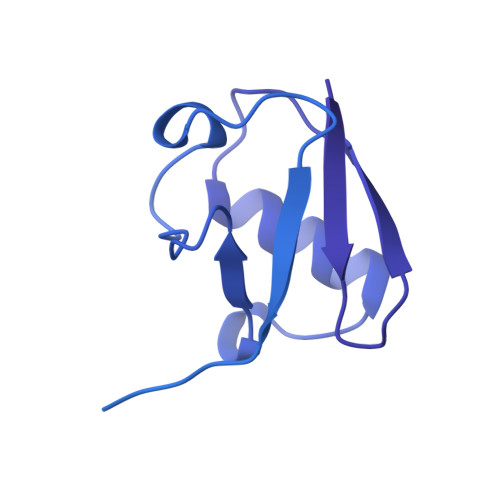
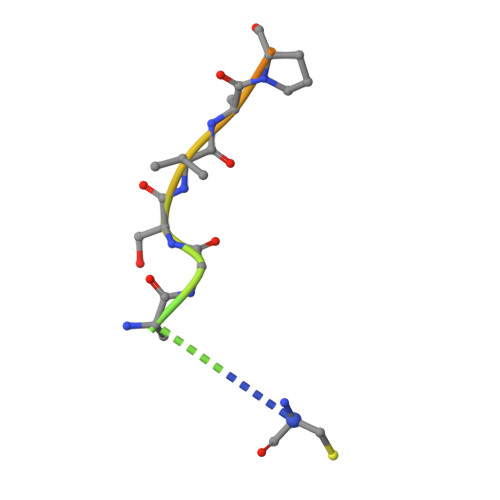
Cullin-RING ligases employ geometrically optimized catalytic partners for substrate targeting.
Li, J., Purser, N., Liwocha, J., Scott, D.C., Byers, H.A., Steigenberger, B., Hill, S., Tripathi-Giesgen, I., Hinkle, T., Hansen, F.M., Prabu, J.R., Radhakrishnan, S.K., Kirkpatrick, D.S., Reichermeier, K.M., Schulman, B.A., Kleiger, G.(2024) Mol Cell 84: 1304
- PubMed: 38382526 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2024.01.022
- Primary Citation Related Structures:
8Q7R, 8R5H - PubMed Abstract:
Cullin-RING ligases (CRLs) ubiquitylate specific substrates selected from other cellular proteins. Substrate discrimination and ubiquitin transferase activity were thought to be strictly separated. Substrates are recognized by substrate receptors, such as Fbox or BCbox proteins. Meanwhile, CRLs employ assorted ubiquitin-carrying enzymes (UCEs, which are a collection of E2 and ARIH-family E3s) specialized for either initial substrate ubiquitylation (priming) or forging poly-ubiquitin chains. We discovered specific human CRL-UCE pairings governing substrate priming. The results reveal pairing of CUL2-based CRLs and UBE2R-family UCEs in cells, essential for efficient PROTAC-induced neo-substrate degradation. Despite UBE2R2's intrinsic programming to catalyze poly-ubiquitylation, CUL2 employs this UCE for geometrically precise PROTAC-dependent ubiquitylation of a neo-substrate and for rapid priming of substrates recruited to diverse receptors. Cryo-EM structures illuminate how CUL2-based CRLs engage UBE2R2 to activate substrate ubiquitylation. Thus, pairing with a specific UCE overcomes E2 catalytic limitations to drive substrate ubiquitylation and targeted protein degradation.
- Department of Chemistry and Biochemistry, University of Nevada, Las Vegas, Las Vegas, NV 89154, USA.
Organizational Affiliation: