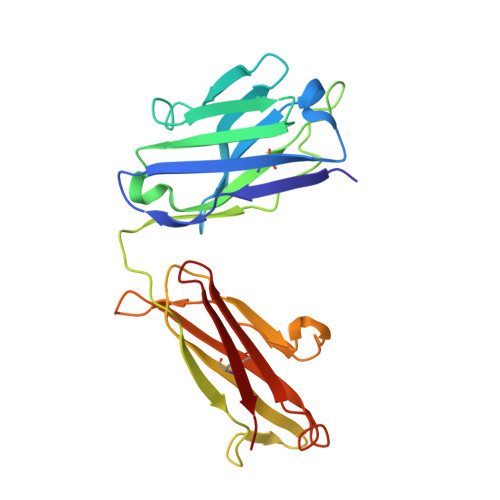
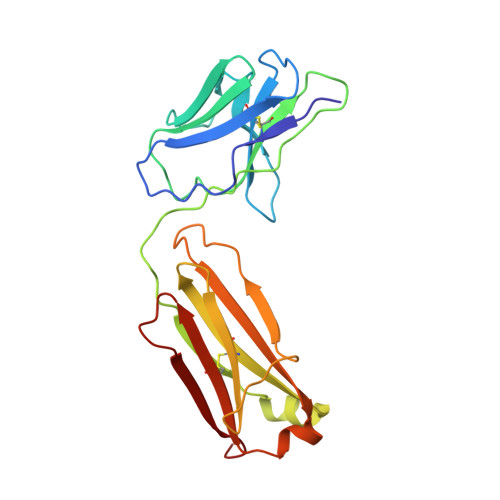
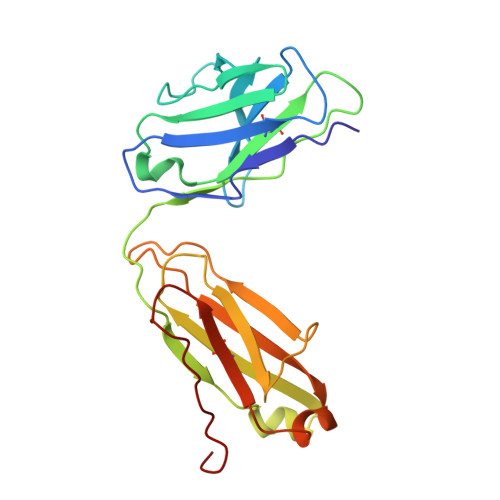
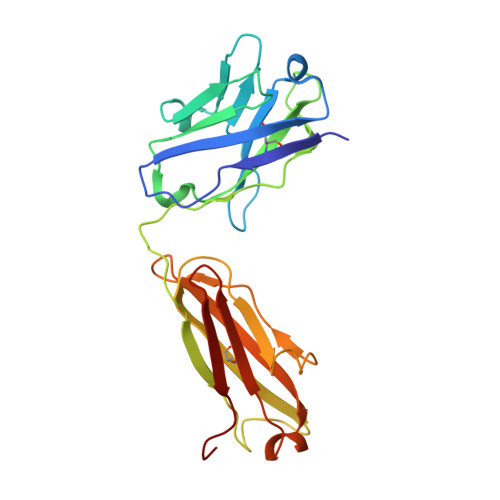
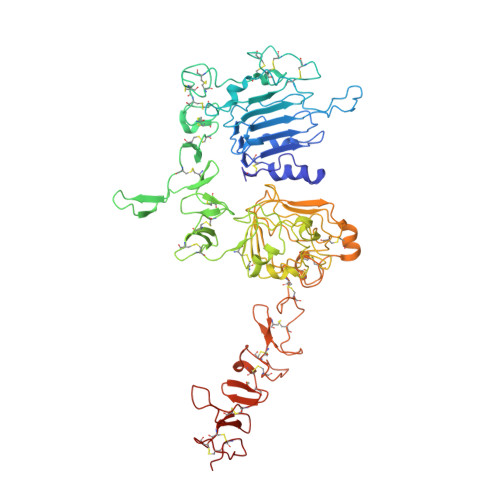
Structure and conformational variability of the HER2-trastuzumab-pertuzumab complex.
Ruedas, R., Vuillemot, R., Tubiana, T., Winter, J.M., Pieri, L., Arteni, A.A., Samson, C., Jonic, S., Mathieu, M., Bressanelli, S.(2024) J Struct Biol 216: 108095-108095
- PubMed: 38723875 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2024.108095
- Primary Citation Related Structures:
8PWH, 8Q6J - PubMed Abstract:
Single particle analysis from cryogenic transmission electron microscopy (cryo-EM) is particularly attractive for complexes for which structure prediction remains intractable, such as antibody-antigen complexes. Here we obtain the detailed structure of a particularly difficult complex between human epidermal growth factor receptor 2 (HER2) and the antigen-binding fragments from two distinct therapeutic antibodies binding to distant parts of the flexible HER2, pertuzumab and trastuzumab (HTP). We highlight the strengths and limitations of current data processing software in dealing with various kinds of heterogeneities, particularly continuous conformational heterogeneity, and in describing the motions that can be extracted from our dataset. Our HTP structure provides a more detailed view than the one previously available for this ternary complex. This allowed us to pinpoint a previously overlooked loop in domain IV that may be involved both in binding of trastuzumab and in HER2 dimerization. This finding may contribute to explain the synergistic anticancer effect of the two antibodies. We further propose that the flexibility of the HTP complex, beyond the difficulties it causes for cryo-EM analysis, actually reflects regulation of HER2 signaling and its inhibition by therapeutic antibodies. Notably we obtain our best data with ultra-thin continuous carbon grids, showing that with current cameras their use to alleviate particle misdistribution is compatible with a protein complex of only 162 kDa. Perhaps most importantly, we provide here a dataset for such a smallish protein complex for further development of software accounting for continuous conformational heterogeneity in cryo-EM images.
- Université Paris-Saclay, CEA, CNRS - Institute for Integrative Biology of the Cell (I2BC), 91198, Gif-sur-Yvette, France; Sanofi, Integrated Drug Discovery, 13, quai Jules Guesde 94403, Vitry-sur-Seine, France.
Organizational Affiliation: