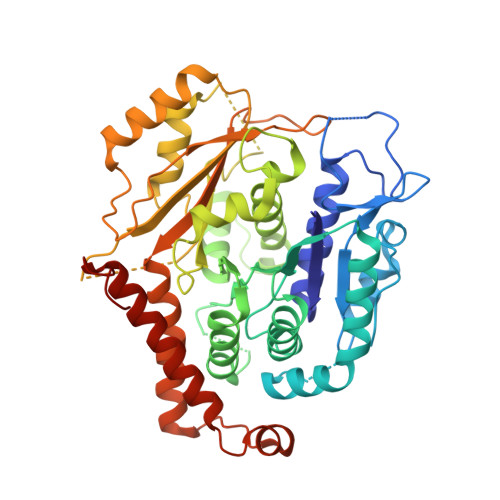
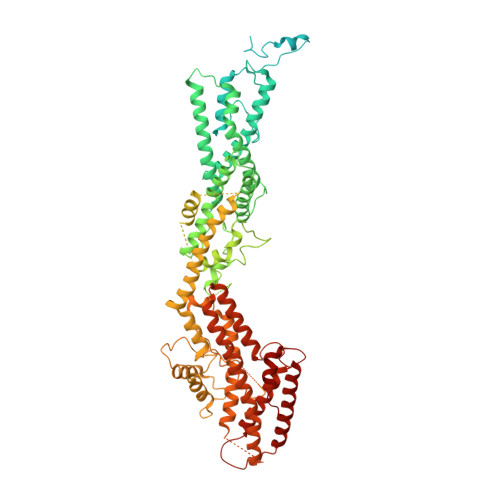
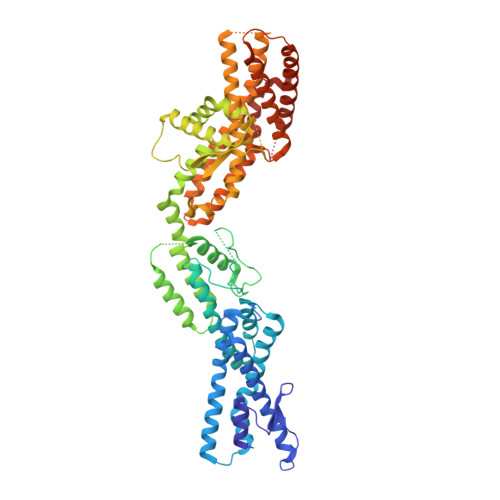
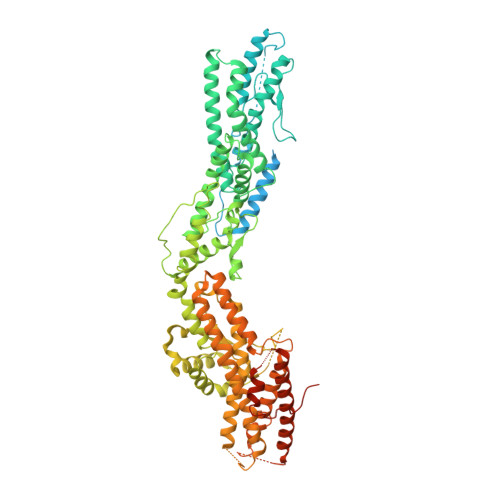
Transition of human gamma-tubulin ring complex into a closed conformation during microtubule nucleation.
Brito, C., Serna, M., Guerra, P., Llorca, O., Surrey, T.(2024) Science 383: 870-876
- PubMed: 38305685 Search on PubMed
- DOI: https://doi.org/10.1126/science.adk6160
- Primary Citation Related Structures:
8Q62 - PubMed Abstract:
Microtubules are essential for intracellular organization and chromosome segregation. They are nucleated by the γ-tubulin ring complex (γTuRC). However, isolated vertebrate γTuRC adopts an open conformation that deviates from the microtubule structure, raising the question of the nucleation mechanism. In this study, we determined cryo-electron microscopy structures of human γTuRC bound to a nascent microtubule. Structural changes of the complex into a closed conformation ensure that γTuRC templates the 13-protofilament microtubules that exist in human cells. Closure is mediated by a latch that interacts with incorporating tubulin, making it part of the closing mechanism. Further rearrangements involve all γTuRC subunits and the removal of the actin-containing luminal bridge. Our proposed mechanism of microtubule nucleation by human γTuRC relies on large-scale structural changes that are likely the target of regulation in cells.
- Centre for Genomic Regulation (CRG), The Barcelona Institute of Science and Technology, Barcelona, Spain.
Organizational Affiliation: