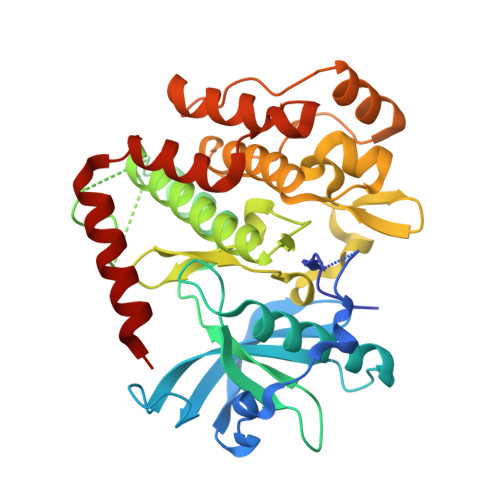
Avapritinib-based SAR studies unveil a binding pocket in KIT and PDGFRA.
Teuber, A., Schulz, T., Fletcher, B.S., Gontla, R., Muhlenberg, T., Zischinsky, M.L., Niggenaber, J., Weisner, J., Kleinbolting, S.B., Lategahn, J., Sievers, S., Muller, M.P., Bauer, S., Rauh, D.(2024) Nat Commun 15: 63-63
- PubMed: 38167404 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-44376-8
- Primary Citation Related Structures:
8PQ9, 8PQA, 8PQB, 8PQC, 8PQD, 8PQE, 8PQF, 8PQG, 8PQH, 8PQI, 8PQJ, 8PQK - PubMed Abstract:
Avapritinib is the only potent and selective inhibitor approved for the treatment of D842V-mutant gastrointestinal stromal tumors (GIST), the most common primary mutation of the platelet-derived growth factor receptor α (PDGFRA). The approval was based on the NAVIGATOR trial, which revealed overall response rates of more than 90%. Despite this transformational activity, patients eventually progress, mostly due to acquired resistance mutations or following discontinuation due to neuro-cognitive side effects. These patients have no therapeutic alternative and face a dismal prognosis. Notable, little is known about this drug's binding mode and its medicinal chemistry development, which is instrumental for the development of the next generation of drugs. Against this background, we solve the crystal structures of avapritinib in complex with wild-type and mutant PDGFRA and stem cell factor receptor (KIT), which provide evidence and understanding of inhibitor binding and lead to the identification of a sub-pocket (Gα-pocket). We utilize this information to design, synthesize and characterize avapritinib derivatives for the determination of key pharmacophoric features to overcome drug resistance and limit potential blood-brain barrier penetration.
- Department of Chemistry and Chemical Biology, TU Dortmund University and Drug Discovery Hub Dortmund (DDHD), Zentrum für Integrierte Wirkstoffforschung (ZIW), Otto-Hahn-Strasse 4a, 44227, Dortmund, Germany.
Organizational Affiliation: