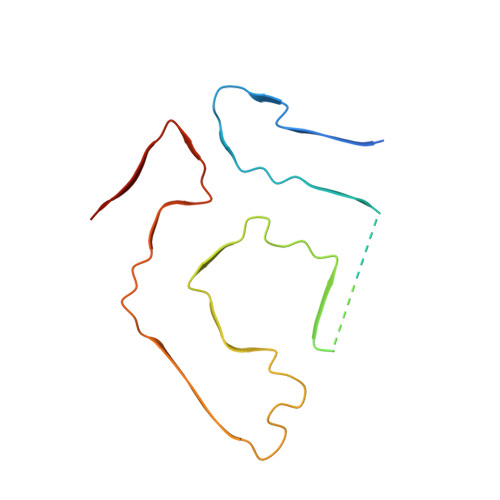
Common transthyretin-derived amyloid fibril structures in patients with hereditary ATTR amyloidosis.
Steinebrei, M., Baur, J., Pradhan, A., Kupfer, N., Wiese, S., Hegenbart, U., Schonland, S.O., Schmidt, M., Fandrich, M.(2023) Nat Commun 14: 7623-7623
- PubMed: 37993462 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-43301-3
- Primary Citation Related Structures:
8PKE, 8PKF, 8PKG - PubMed Abstract:
Systemic ATTR amyloidosis is an increasingly important protein misfolding disease that is provoked by the formation of amyloid fibrils from transthyretin protein. The pathological and clinical disease manifestations and the number of pathogenic mutational changes in transthyretin are highly diverse, raising the question whether the different mutations may lead to different fibril morphologies. Using cryo-electron microscopy, however, we show here that the fibril structure is remarkably similar in patients that are affected by different mutations. Our data suggest that the circumstances under which these fibrils are formed and deposited inside the body - and not only the fibril morphology - are crucial for defining the phenotypic variability in many patients.
- Institute of Protein Biochemistry, Ulm University, Helmholtzstrasse 8/1, Ulm, D-89081, Germany. maximilian.steinebrei@uni-ulm.de.
Organizational Affiliation: