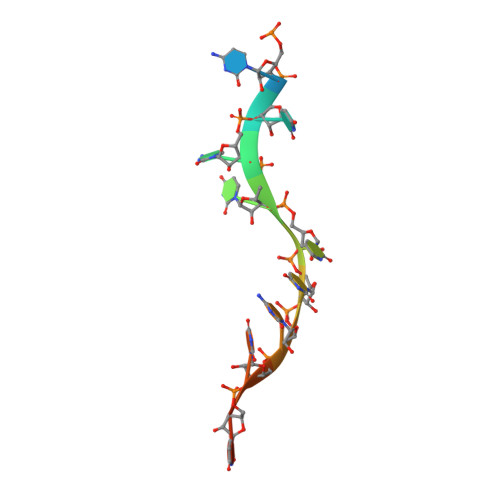
Structural basis of RNA-induced autoregulation of the DExH-type RNA helicase maleless.
Jagtap, P.K.A., Muller, M., Kiss, A.E., Thomae, A.W., Lapouge, K., Beck, M., Becker, P.B., Hennig, J.(2023) Mol Cell 83: 4318-4333.e10
- PubMed: 37989319 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2023.10.026
- Primary Citation Related Structures:
8B9G, 8B9I, 8B9J, 8B9K, 8B9L, 8PJB, 8PJJ - PubMed Abstract:
RNA unwinding by DExH-type helicases underlies most RNA metabolism and function. It remains unresolved if and how the basic unwinding reaction of helicases is regulated by auxiliary domains. We explored the interplay between the RecA and auxiliary domains of the RNA helicase maleless (MLE) from Drosophila using structural and functional studies. We discovered that MLE exists in a dsRNA-bound open conformation and that the auxiliary dsRBD2 domain aligns the substrate RNA with the accessible helicase tunnel. In an ATP-dependent manner, dsRBD2 associates with the helicase module, leading to tunnel closure around ssRNA. Furthermore, our structures provide a rationale for blunt-ended dsRNA unwinding and 3'-5' translocation by MLE. Structure-based MLE mutations confirm the functional relevance of our model for RNA unwinding. Our findings contribute to our understanding of the fundamental mechanics of auxiliary domains in DExH helicase MLE, which serves as a model for its human ortholog and potential therapeutic target, DHX9/RHA.
- Structural and Computational Biology Unit, EMBL Heidelberg, Meyerhofstraße 1, 69117 Heidelberg, Germany; Chair of Biochemistry IV, Biophysical Chemistry, University of Bayreuth, Bayreuth, Germany. Electronic address: pravin.jagtap@embl.de.
Organizational Affiliation: