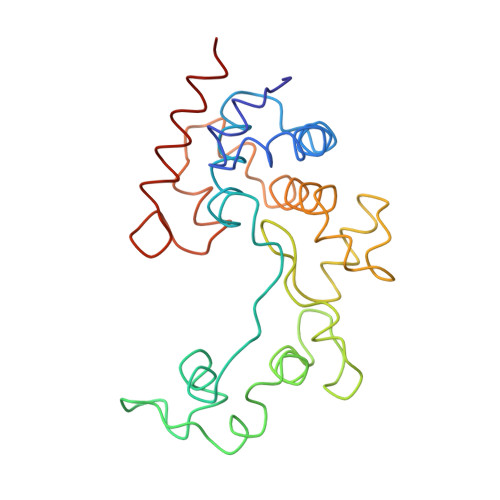
Structural and Biochemical Characterization of a New Phage-Encoded Muramidase, KTN6 Gp46.
Sanz-Gaitero, M., De Maesschalck, V., Patel, A., Longin, H., Van Noort, V., Rodriguez-Rubio, L., van Ryne, M., Danis-Wlodarczyk, K., Drulis-Kawa, Z., Mesnage, S., van Raaij, M., Lavigne, R.(2024) Phage (New Rochelle) 5: 53-62
- PubMed: 39119210 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1089/phage.2023.0040
- Primary Citation Related Structures:
8P8E - PubMed Abstract:
Endolysins are phage-encoded lytic enzymes that degrade bacterial peptidoglycan at the end of phage lytic cycles to release new phage particles. These enzymes are being explored as an alternative to small-molecule antibiotics. The crystal structure of KTN6 Gp46 was determined and compared with a ColabFold model. Cleavage specificity was examined using a peptidoglycan digest and reversed-phase high-performance liquid chromatography coupled to mass spectrometry (HPLC/MS). The structure of KTN6 Gp46 could be determined at 1.4 Å resolution, and key differences in loops of the putative peptidoglycan binding domain were identified in comparison with its closest known homologue, the endolysin of phage SPN1S. Reversed-phase HPLC/MS analysis of the reaction products following peptidoglycan digestion confirmed the muramidase activity of Gp46, consistent with structural predictions. These insights into the structure and function of endolysins further expand the toolbox for endolysin engineering and explore their potential in enzyme-based antibacterial design strategies.
- Centro Nacional de Biotecnologia, Consejo Superior de Investigaciones Cientificas, Madrid, Spain.
Organizational Affiliation: