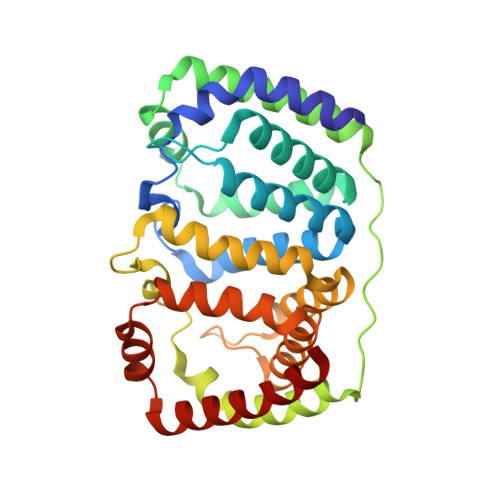
Structural and spectroscopic characterization of the peridinin-chlorophyll a-protein (PCP) complex from Heterocapsa pygmaea (HPPCP).
Schulte, T., Magdaong, N.C.M., Di Valentin, M., Agostini, A., Tait, C.E., Niedzwiedzki, D.M., Carbonera, D., Hofmann, E.(2024) Biochim Biophys Acta Bioenerg 1866: 149510-149510
- PubMed: 39321862 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbabio.2024.149510
- Primary Citation Related Structures:
8OV5, 8OW6 - PubMed Abstract:
Light harvesting proteins are optimized to efficiently collect and transfer light energy for photosynthesis. In eukaryotic dinoflagellates these complexes utilize chlorophylls and a special carotenoid, peridinin, and arrange them for efficient excitation energy transfer. At the same time, the carotenoids protect the system by quenching harmful chlorophyll triplet states. Here we use advanced spectroscopic techniques and X-ray structure analysis to investigate excitation energy transfer processes in the major soluble antenna, the peridinin chlorophyll a protein (PCP) from the free living dinoflagellate Heterocapsa pygmaea. We determined the 3D-structure of this complex at high resolution (1.2 Å). For better comparison, we improved the reference structure of this protein from Amphidinium carterae to a resolution of 1.15 Å. We then used fs and ns time-resolved absorption spectroscopy to study the mechanisms of light harvesting, but also of the photoprotective quenching of the chlorophyll triplet state. The photoprotection site was further characterized by Electron Spin Echo Envelope Modulation (ESEEM) spectroscopy to yield information on water molecules involved in triplet-triplet energy transfer. Similar to other PCP complexes, excitation energy transfer from peridinin to chlorophyll is found to be very efficient, with transfer times in the range of 1.6-2.1 ps. One of the four carotenoids, the peridinin 614, is well positioned to quench the chlorophyll triplet state with high efficiency and transfer times in the range of tens of picoseconds. Our structural and dynamic data further support, that the intrinsic water molecule coordinating the chlorophyll Mg ion plays an essential role in photoprotection.
- Protein Crystallography, Faculty of Biology and Biotechnology, Ruhr University Bochum, 44801 Bochum, Germany; Department of Biochemistry and Biophysics, National Bioinformatics Infrastructure Sweden, Science for Life Laboratory, Stockholm University, 17121 Solna, Sweden.
Organizational Affiliation: